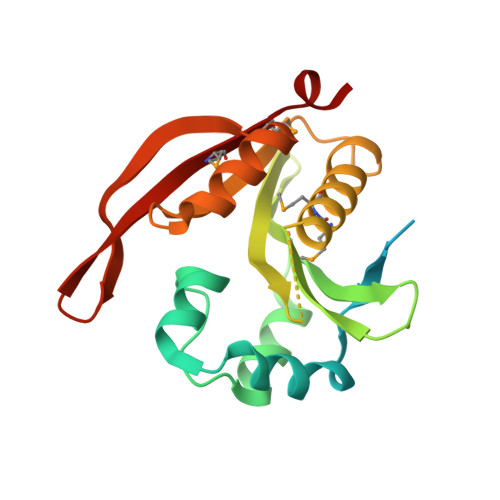
Functional and structural characterisation of RimL from Bacillus cereus, a new N alpha-acetyltransferase of ribosomal proteins that was wrongly assigned as an aminoglycosyltransferase.
Leonardo Silvestre, H., Asensio, J.L., Blundell, T.L., Bastida, A., Bolanos-Garcia, V.M.(2024) Int J Biol Macromol 263: 130348-130348
- PubMed: 38395274 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2024.130348
- Primary Citation Related Structures:
6ERD, 8Q6U - PubMed Abstract:
Enzymes of the GNAT (GCN5-relate N-acetyltransferases) superfamily are important regulators of cell growth and development. They are functionally diverse and share low amino acid sequence identity, making functional annotation difficult. In this study, we report the function and structure of a new ribosomal enzyme, N α -acetyl transferase from Bacillus cereus (RimL BC ), a protein that was previously wrongly annotated as an aminoglycosyltransferase. Firstly, extensive comparative amino acid sequence analyses suggested RimL BC belongs to a cluster of proteins mediating acetylation of the ribosomal protein L7/L12. To assess if this was the case, several well established substrates of aminoglycosyltransferases were screened. The results of these studies did not support an aminoglycoside acetylating function for RimL BC . To gain further insight into RimL BC biological role, a series of studies that included MALDI-TOF, isothermal titration calorimetry, NMR, X-ray protein crystallography, and site-directed mutagenesis confirmed RimL BC affinity for Acetyl-CoA and that the ribosomal protein L7/L12 is a substrate of RimL BC . Last, we advance a mechanistic model of RimL BC mode of recognition of its protein substrates. Taken together, our studies confirmed RimL BC as a new ribosomal N α -acetyltransferase and provide structural and functional insights into substrate recognition by N α -acetyltransferases and protein acetylation in bacteria.
- Department of Biochemistry, University of Cambridge, 80 Tennis Court Road, Cambridge CB2 1GA, United Kingdom.
Organizational Affiliation: