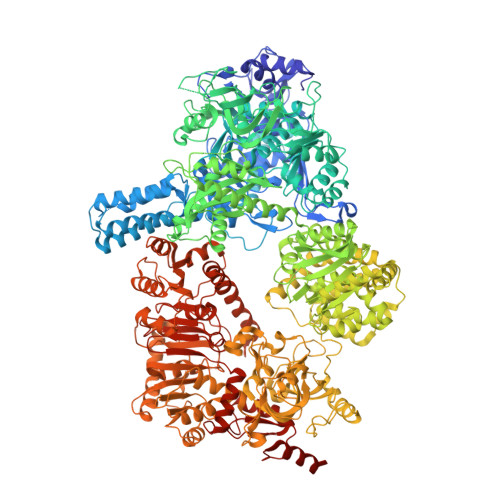
The multicatalytic compartment of propionyl-CoA synthase sequesters a toxic metabolite.
Bernhardsgrutter, I., Vogeli, B., Wagner, T., Peter, D.M., Cortina, N.S., Kahnt, J., Bange, G., Engilberge, S., Girard, E., Riobe, F., Maury, O., Shima, S., Zarzycki, J., Erb, T.J.(2018) Nat Chem Biol 14: 1127-1132
- PubMed: 30374166
- DOI: https://doi.org/10.1038/s41589-018-0153-x
- Primary Citation Related Structures:
6EQO - PubMed Abstract:
Cells must cope with toxic or reactive intermediates formed during metabolism. One coping strategy is to sequester reactions that produce such intermediates within specialized compartments or tunnels connecting different active sites. Here, we show that propionyl-CoA synthase (PCS), an ∼ 400-kDa homodimer, three-domain fusion protein and the key enzyme of the 3-hydroxypropionate bi-cycle for CO 2 fixation, sequesters its reactive intermediate acrylyl-CoA. Structural analysis showed that PCS forms a multicatalytic reaction chamber. Kinetic analysis suggested that access to the reaction chamber and catalysis are synchronized by interdomain communication. The reaction chamber of PCS features three active sites and has a volume of only 33 nm 3 . As one of the smallest multireaction chambers described in biology, PCS may inspire the engineering of a new class of dynamically regulated nanoreactors.
- Department of Biochemistry and Synthetic Metabolism, Max Planck Institute for Terrestrial Microbiology, Marburg, Germany.
Organizational Affiliation: