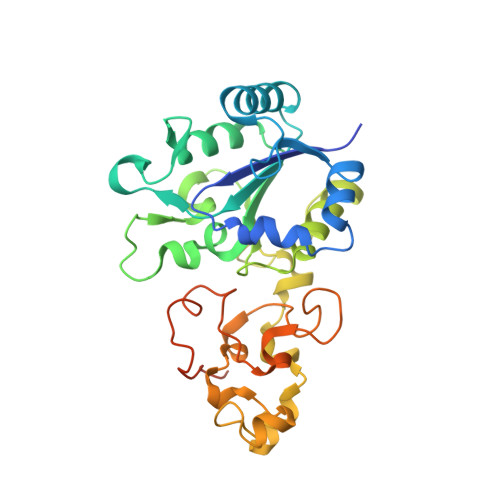
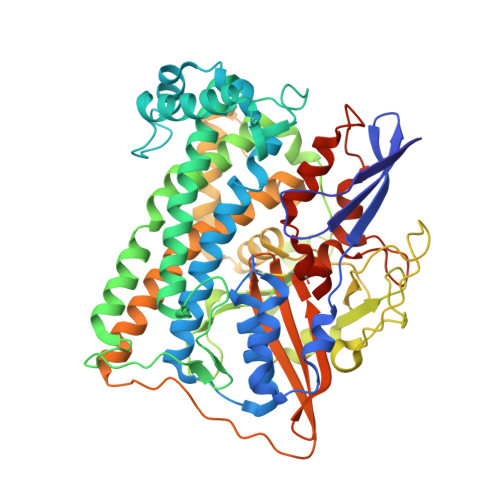
The structure of hydrogenase-2 fromEscherichia coli: implications for H2-driven proton pumping.
Beaton, S.E., Evans, R.M., Finney, A.J., Lamont, C.M., Armstrong, F.A., Sargent, F., Carr, S.B.(2018) Biochem J 475: 1353-1370
- PubMed: 29555844 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BCJ20180053
- Primary Citation Related Structures:
6EHQ, 6EHS, 6EN9 - PubMed Abstract:
Under anaerobic conditions, Escherichia coli is able to metabolize molecular hydrogen via the action of several [NiFe]-hydrogenase enzymes. Hydrogenase-2, which is typically present in cells at low levels during anaerobic respiration, is a periplasmic-facing membrane-bound complex that functions as a proton pump to convert energy from hydrogen (H 2 ) oxidation into a proton gradient; consequently, its structure is of great interest. Empirically, the complex consists of a tightly bound core catalytic module, comprising large (HybC) and small (HybO) subunits, which is attached to an Fe-S protein (HybA) and an integral membrane protein (HybB). To date, efforts to gain a more detailed picture have been thwarted by low native expression levels of Hydrogenase-2 and the labile interaction between HybOC and HybA/HybB subunits. In the present paper, we describe a new overexpression system that has facilitated the determination of high-resolution crystal structures of HybOC and, hence, a prediction of the quaternary structure of the HybOCAB complex.
- Department of Chemistry, Inorganic Chemistry Laboratory, University of Oxford, Oxford OX1 3QR, U.K.
Organizational Affiliation: