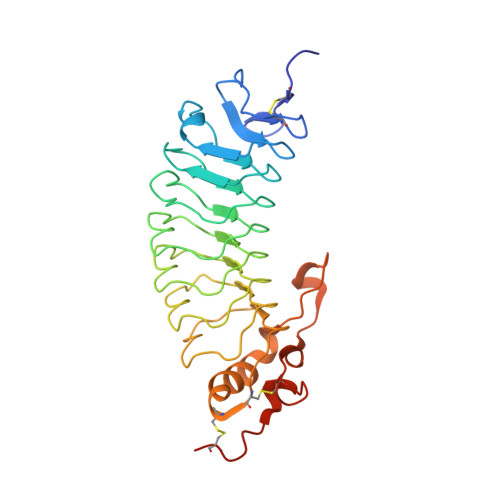
Structural basis of the leukocyte integrin Mac-1 I-domain interactions with the platelet glycoprotein Ib.
Morgan, J., Saleem, M., Ng, R., Armstrong, C., Wong, S.S., Caulton, S.G., Fickling, A., Williams, H.E.L., Munday, A.D., Lopez, J.A., Searle, M.S., Emsley, J.(2019) Blood Adv 3: 1450-1459
- PubMed: 31053572 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1182/bloodadvances.2018027011
- Primary Citation Related Structures:
6EJX - PubMed Abstract:
Cell-surface receptor interactions between leukocyte integrin macrophage-1 antigen (Mac-1, also known as CR3, αMβ2, CD11b/CD18) and platelet glycoprotein Ibα (GPIbα) are critical to vascular inflammation. To define the key residues at the binding interface, we used nuclear magnetic resonance (NMR) to assign the spectra of the mouse Mac-1 I-domain and mapped the residues contacting the mouse GPIbα N-terminal domain (GPIbαN) to the locality of the integrin metal ion-dependant adhesion site (MIDAS) surface. We next determined the crystal structures of the mouse GPIbαN and Mac-1 I-domain to 2 Å and 2.5 Å resolution, respectively. The mouse Mac-1 I-domain crystal structure reveals an active conformation that is stabilized by a crystal contact from the α7-helix with a glutamate side chain completing the octahedral coordination sphere of the MIDAS Mg 2+ ion. The amino acid sequence of the α7-helix and disposition of the glutamic acid matches the C-terminal capping region α-helix of GPIbα effectively acting as a ligand mimetic. Using these crystal structures in combination with NMR measurements and docking analysis, we developed a model whereby an acidic residue from the GPIbα leucine-rich repeat (LRR) capping α-helix coordinates directly to the Mac-1 MIDAS Mg 2+ ion. The Mac-1:GPIbαN complex involves additional interactions consolidated by an elongated pocket flanking the GPIbαN LRR capping α-helix. The GPIbαN α-helix has an HxxxE motif, which is equivalent by homology to RxxxD from the human GPIbαN. Subsequent mutagenesis of residues at this interface, coupled with surface plasmon resonance studies, confirmed the importance of GPIbαN residues H218, E222, and the Mac-1 MIDAS residue T209 to formation of the complex.
- School of Chemistry and.
Organizational Affiliation: