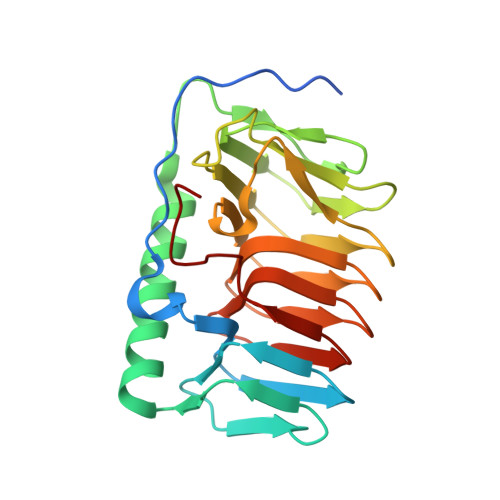
Structure of a bacterial ice binding protein with two faces of interaction with ice.
Mangiagalli, M., Sarusi, G., Kaleda, A., Bar Dolev, M., Nardone, V., Vena, V.F., Braslavsky, I., Lotti, M., Nardini, M.(2018) FEBS J 285: 1653-1666
- PubMed: 29533528 Search on PubMed
- DOI: https://doi.org/10.1111/febs.14434
- Primary Citation Related Structures:
6EIO - PubMed Abstract:
Ice-binding proteins (IBPs) contribute to the survival of many living beings at subzero temperature by controlling the formation and growth of ice crystals. This work investigates the structural basis of the ice-binding properties of EfcIBP, obtained from Antarctic bacteria. EfcIBP is endowed with a unique combination of thermal hysteresis and ice recrystallization inhibition activity. The three-dimensional structure, solved at 0.84 Å resolution, shows that EfcIBP belongs to the IBP-1 fold family, and is organized in a right-handed β-solenoid with a triangular cross-section that forms three protein surfaces, named A, B, and C faces. However, EfcIBP diverges from other IBP-1 fold proteins in relevant structural features including the lack of a 'capping' region on top of the β-solenoid, and in the sequence and organization of the regions exposed to ice that, in EfcIBP, reveal the presence of threonine-rich ice-binding motifs. Docking experiments and site-directed mutagenesis pinpoint that EfcIBP binds ice crystals not only via its B face, as common to other IBPs, but also via ice-binding sites on the C face. Coordinates and structure factors have been deposited in the Protein Data Bank under accession number 6EIO.
- Department of Biotechnology and Biosciences, University of Milano-Bicocca, Italy.
Organizational Affiliation: