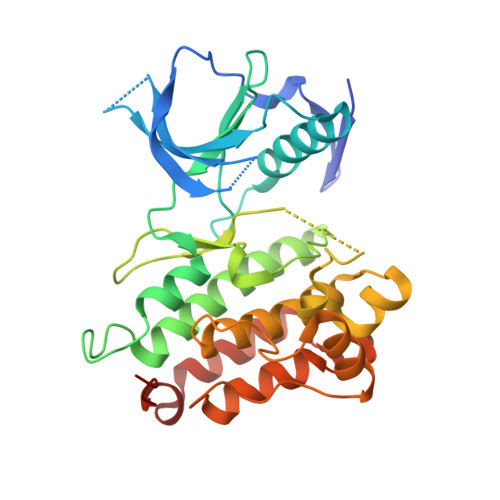
Discovery of Potent, Selective, and Brain-Penetrant 1 H-Pyrazol-5-yl-1 H-pyrrolo[2,3- b]pyridines as Anaplastic Lymphoma Kinase (ALK) Inhibitors.
Fushimi, M., Fujimori, I., Wakabayashi, T., Hasui, T., Kawakita, Y., Imamura, K., Kato, T., Murakami, M., Ishii, T., Kikko, Y., Kasahara, M., Nakatani, A., Hiura, Y., Miyamoto, M., Saikatendu, K., Zou, H., Lane, S.W., Lawson, J.D., Imoto, H.(2019) J Med Chem 62: 4915-4935
- PubMed: 31009559 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.8b01630
- Primary Citation Related Structures:
6E0R, 6EBW, 6EDL - PubMed Abstract:
Anaplastic lymphoma kinase (ALK), a member of the receptor tyrosine kinase family, is predominantly expressed in the brain and implicated in neuronal development and cognition. However, the detailed function of ALK in the central nervous system (CNS) is still unclear. To elucidate the role of ALK in the CNS, it was necessary to discover a potent, selective, and brain-penetrant ALK inhibitor. Scaffold hopping and lead optimization of N-(2,4-difluorobenzyl)-3-(1 H-pyrazol-5-yl)imidazo[1,2- b]pyridazin-6-amine 1 guided by a cocrystal structure of compound 1 bound to ALK resulted in the identification of (6-(1-(5-fluoropyridin-2-yl)ethoxy)-1-(5-methyl-1 H-pyrazol-3-yl)-1 H-pyrrolo[2,3- b]pyridin-3-yl)((2 S)-2-methylmorpholin-4-yl)methanone 13 as a highly potent, selective, and brain-penetrable compound. Intraperitoneal administration of compound 13 significantly decreased the phosphorylated-ALK (p-ALK) levels in the hippocampus and prefrontal cortex in the mouse brain. These results suggest that compound 13 could serve as a useful chemical probe to elucidate the mechanism of ALK-mediated brain functions and the therapeutic potential of ALK inhibition.
- Neuroscience Drug Discovery Unit, Research , Takeda Pharmaceutical Company Limited , 26-1, Muraoka-Higashi 2-chome, Fujisawa , Kanagawa 251-8555 , Japan.
Organizational Affiliation: