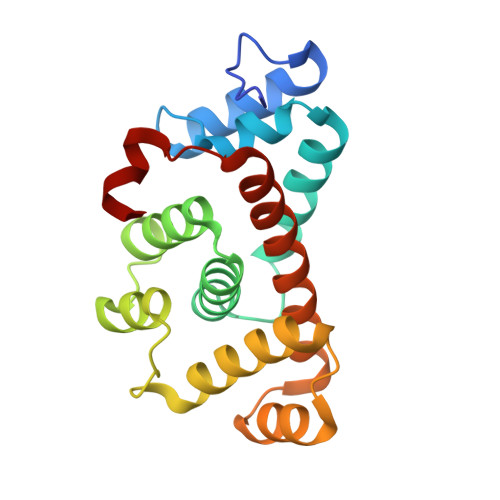
Structure-function analyses of alkylhydroperoxidase D fromStreptococcus pneumoniaereveal an unusual three-cysteine active site architecture.
Meng, Y., Sheen, C.R., Magon, N.J., Hampton, M.B., Dobson, R.C.J.(2020) J Biological Chem 295: 2984-2999
- PubMed: 31974167 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.012226
- Primary Citation Related Structures:
6E8L - PubMed Abstract:
During aerobic growth, the Gram-positive facultative anaerobe and opportunistic human pathogen Streptococcus pneumoniae generates large amounts of hydrogen peroxide that can accumulate to millimolar concentrations. The mechanism by which this catalase-negative bacterium can withstand endogenous hydrogen peroxide is incompletely understood. The enzyme alkylhydroperoxidase D (AhpD) has been shown to contribute to pneumococcal virulence and oxidative stress responses in vivo We demonstrate here that Sp AhpD exhibits weak thiol-dependent peroxidase activity and, unlike the previously reported Mycobacterium tuberculosis AhpC/D system, Sp AhpD does not mediate electron transfer to Sp AhpC. A 2.3-Å resolution crystal structure revealed several unusual structural features, including a three-cysteine active site architecture that is buried in a deep pocket, in contrast to the two-cysteine active site found in other AhpD enzymes. All single-cysteine Sp AhpD variants remained partially active, and LC-MS/MS analyses revealed that the third cysteine, Cys-163, formed disulfide bonds with either of two cysteines in the canonical Cys-78- X-X -Cys-81 motif. We observed that Sp AhpD formed a dimeric quaternary structure both in the crystal and in solution, and that the highly conserved Asn-76 of the AhpD core motif is important for Sp AhpD folding. In summary, Sp AhpD is a weak peroxidase and does not transfer electrons to AhpC, and therefore does not fit existing models of bacterial AhpD antioxidant defense mechanisms. We propose that it is unlikely that Sp AhpD removes peroxides either directly or via AhpC, and that Sp AhpD cysteine oxidation may act as a redox switch or mediate electron transfer with other thiol proteins.
- Biomolecular Interaction Centre and School of Biological Sciences, University of Canterbury, Christchurch 8041, New Zealand.
Organizational Affiliation: