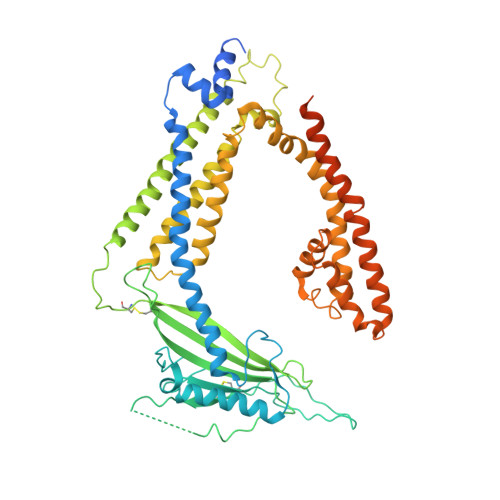
Structural basis for PtdInsP2-mediated human TRPML1 regulation.
Fine, M., Schmiege, P., Li, X.(2018) Nat Commun 9: 4192-4192
- PubMed: 30305615 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-06493-7
- Primary Citation Related Structures:
6E7P, 6E7Y, 6E7Z - PubMed Abstract:
Transient receptor potential mucolipin 1 (TRPML1), a lysosomal channel, maintains the low pH and calcium levels for lysosomal function. Several small molecules modulate TRPML1 activity. ML-SA1, a synthetic agonist, binds to the pore region and phosphatidylinositol-3,5-bisphosphate (PtdIns(3,5)P 2 ), a natural lipid, stimulates channel activity to a lesser extent than ML-SA1; moreover, PtdIns(4,5)P 2 , another natural lipid, prevents TRPML1-mediated calcium release. Notably, PtdIns(3,5)P 2 and ML-SA1 cooperate further increasing calcium efflux. Here we report the structures of human TRPML1 at pH 5.0 with PtdIns(3,5)P 2 , PtdIns(4,5)P 2 , or ML-SA1 and PtdIns(3,5)P 2 , revealing a unique lipid-binding site. PtdIns(3,5)P 2 and PtdIns(4,5)P 2 bind to the extended helices of S1, S2, and S3. The phosphate group of PtdIns(3,5)P 2 induces Y355 to form a π-cation interaction with R403, moving the S4-S5 linker, thus allosterically activating the channel. Our structures and electrophysiological characterizations reveal an allosteric site and provide molecular insight into how lipids regulate TRP channels.
- Department of Physiology, University of Texas Southwestern Medical Center, Dallas, TX, 75390, USA.
Organizational Affiliation: