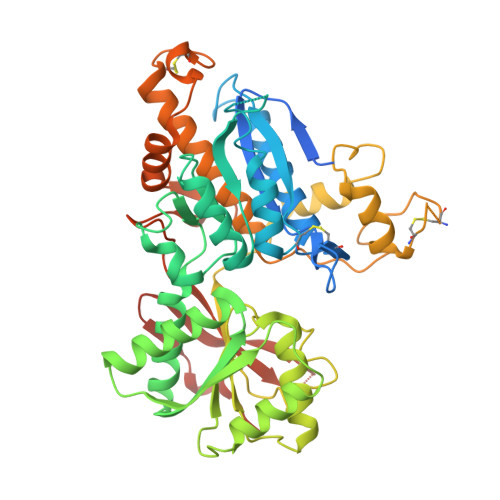
Structural Basis for ( S)-3,4-Dicarboxyphenylglycine (DCPG) As a Potent and Subtype Selective Agonist of the mGlu8Receptor.
Chen, Q., Ho, J.D., Ashok, S., Vargas, M.C., Wang, J., Atwell, S., Bures, M., Schkeryantz, J.M., Monn, J.A., Hao, J.(2018) J Med Chem 61: 10040-10052
- PubMed: 30365309 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.8b01120
- Primary Citation Related Structures:
6E5V - PubMed Abstract:
( S)-3,4-Dicarboxyphenylglycine (DCPG) was first reported in 2001 as a potent orthosteric agonist with high subtype selectivity for the mGlu 8 receptor, but the structural basis for its high selectivity is not well understood. We have solved a cocrystal structure of recombinant human mGlu 8 amino terminal domain (ATD) protein bound to ( S)-DCPG, which possesses the largest lobe opening angle observed to date among known agonist-bound mGlu ATD crystal structures. The binding conformation of ( S)-DCPG observed in the crystal structure is significantly different from that in the homology model built from an l-glutamate-bound rat mGlu 1 ATD crystal structure, which has a smaller lobe opening angle. This highlights the importance of considering various lobe opening angles when modeling mGlu ATD-ligand complex. New homology models of other mGlu receptors based on the ( S)-DCPG-bound mGlu 8 ATD crystal structure were explored to rationalize ( S)-DCPG's high mGlu 8 receptor subtype selectivity.