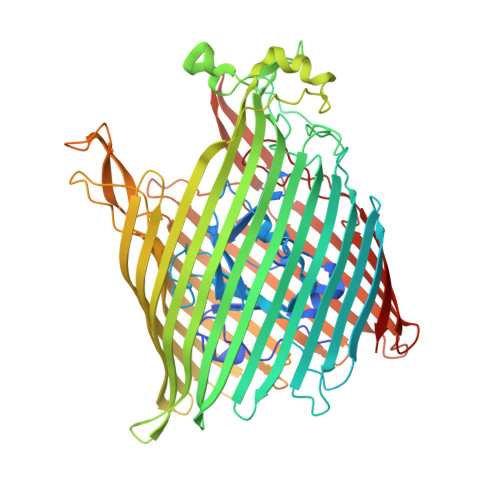
Determination of the molecular basis for coprogen import by Gram-negative bacteria.
Grinter, R., Lithgow, T.(2019) IUCrJ 6: 401-411
- PubMed: 31098021 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2052252519002926
- Primary Citation Related Structures:
6E4V - PubMed Abstract:
In order to survive in mixed microbial communities, some species of fungi secrete coprogens, siderophores that facilitate capture of the scarce nutrient iron. The TonB-dependent transporter FhuE is integrated in the outer membrane of Gram-negative bacteria and has been reported to scavenge these fungally produced coprogens. In this work, an Escherichia coli strain was engineered that is dependent solely on FhuE for its access to siderophore-sequestered iron. Using this tool, it is shown that while FhuE is highly active in the import of coprogens, it has some level of promiscuity, acting as a low-affinity transporter for related siderophores. The crystal structure of FhuE in complex with coprogen was determined, providing a structural basis to explain this selective promiscuity. The structural data, in combination with functional analysis, presented in this work show that FhuE has evolved to specifically engage with planar siderophores. A potential evolutionary driver, and a critical consequence of this selectivity, is that it allows FhuE to exclude antibiotics that mimic nonplanar hydroxamate siderophores: these toxic molecules could otherwise cross the outer membrane barrier through a Trojan horse mechanism.
- Infection and Immunity Program, Biomedicine Discovery Institute and Department of Microbiology, Monash University, Monash, Victoria 3800, Australia.
Organizational Affiliation: