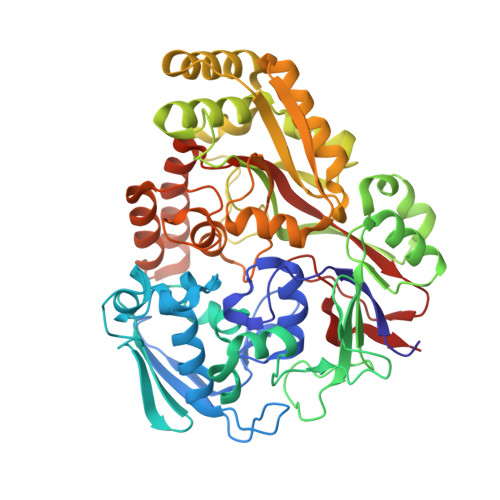
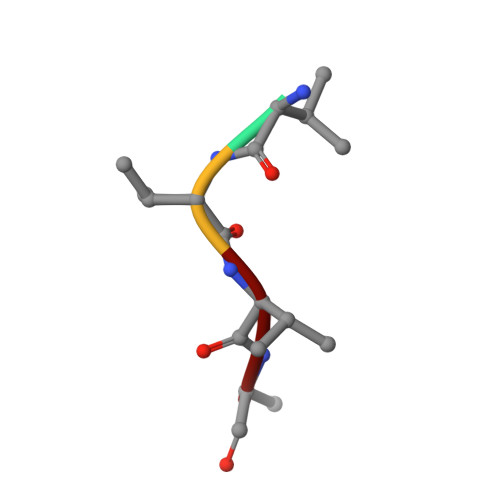
Heme and hemoglobin utilization by Mycobacterium tuberculosis.
Mitra, A., Ko, Y.H., Cingolani, G., Niederweis, M.(2019) Nat Commun 10: 4260-4260
- PubMed: 31534126 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-12109-5
- Primary Citation Related Structures:
6E3D, 6E4D - PubMed Abstract:
Iron is essential for growth of Mycobacterium tuberculosis (Mtb), but most iron in the human body is stored in heme within hemoglobin. Here, we demonstrate that the substrate-binding protein DppA of the inner membrane Dpp transporter is required for heme and hemoglobin utilization by Mtb. The 1.27 Å crystal structure of DppA shows a tetrapeptide bound in the protein core and a large solvent-exposed crevice for heme binding. Mutation of arginine 179 in this cleft eliminates heme binding to DppA and prevents heme utilization by Mtb. The outer membrane proteins PPE36 and PPE62 are also required for heme and hemoglobin utilization, indicating that these pathways converge at the cell surface of Mtb. Albumin, the most abundant blood protein, binds heme specifically and bypasses the requirements for PPE36, PPE62 and Dpp. Thus, our study reveals albumin-dependent and -independent heme uptake pathways, highlighting the importance of iron acquisition from heme for Mtb.
- Department of Microbiology, University of Alabama at Birmingham, 845 19th Street South, Birmingham, AL, 35294, USA.
Organizational Affiliation: