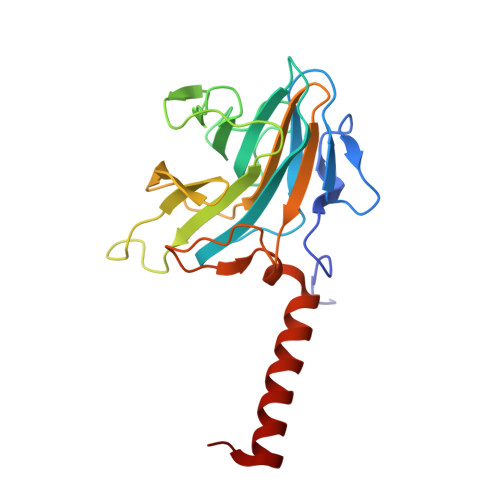
Structural Analysis of the Ash2L/Dpy-30 Complex Reveals a Heterogeneity in H3K4 Methylation.
Haddad, J.F., Yang, Y., Takahashi, Y.H., Joshi, M., Chaudhary, N., Woodfin, A.R., Benyoucef, A., Yeung, S., Brunzelle, J.S., Skiniotis, G., Brand, M., Shilatifard, A., Couture, J.F.(2018) Structure 26: 1594
- PubMed: 30270175 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2018.08.004
- Primary Citation Related Structures:
6E2H - PubMed Abstract:
Dpy-30 is a regulatory subunit controlling the histone methyltransferase activity of the KMT2 enzymes in vivo. Paradoxically, in vitro methyltransferase assays revealed that Dpy-30 only modestly participates in the positive heterotypic allosteric regulation of these methyltransferases. Detailed genome-wide, molecular and structural studies reveal that an extensive network of interactions taking place at the interface between Dpy-30 and Ash2L are critical for the correct placement, genome-wide, of H3K4me2 and H3K4me3 but marginally contribute to the methyltransferase activity of KMT2 enzymes in vitro. Moreover, we show that H3K4me2 peaks persisting following the loss of Dpy-30 are found in regions of highly transcribed genes, highlighting an interplay between Complex of Proteins Associated with SET1 (COMPASS) kinetics and the cycling of RNA polymerase to control H3K4 methylation. Overall, our data suggest that Dpy-30 couples its modest positive heterotypic allosteric regulation of KMT2 methyltransferase activity with its ability to help the positioning of SET1/COMPASS to control epigenetic signaling.
- Ottawa Institute of Systems Biology, Department of Biochemistry, Microbiology and Immunology, University of Ottawa, 451 Smyth Road, Roger Guindon Hall, Ottawa, ON K1H 8M5, Canada.
Organizational Affiliation: