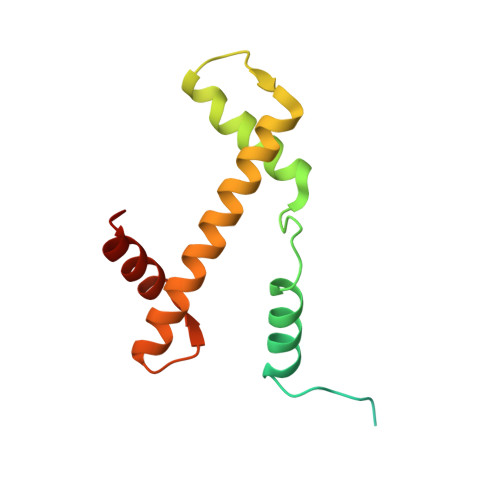
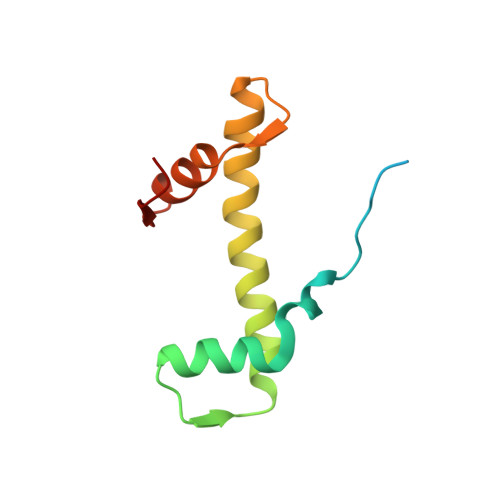
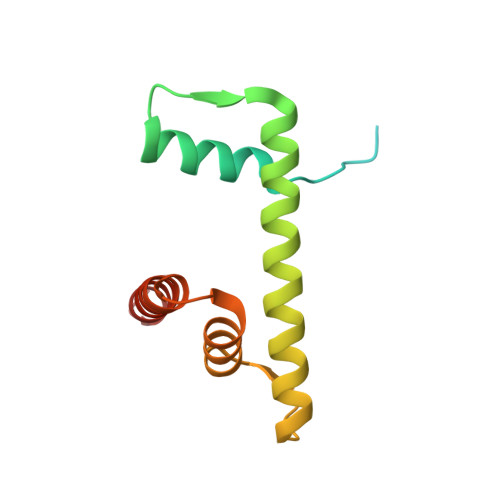
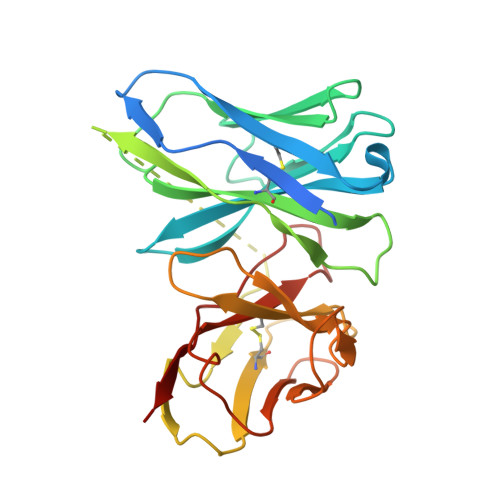
Atomic resolution cryo-EM structure of a native-like CENP-A nucleosome aided by an antibody fragment.
Zhou, B.R., Yadav, K.N.S., Borgnia, M., Hong, J., Cao, B., Olins, A.L., Olins, D.E., Bai, Y., Zhang, P.(2019) Nat Commun 10: 2301-2301
- PubMed: 31127102 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-10247-4
- Primary Citation Related Structures:
6DZT, 6E0C, 6E0P, 6O1D - PubMed Abstract:
Genomic DNA in eukaryotes is organized into chromatin through association with core histones to form nucleosomes, each distinguished by their DNA sequences and histone variants. Here, we used a single-chain antibody fragment (scFv) derived from the anti-nucleosome antibody mAb PL2-6 to stabilize human CENP-A nucleosome containing a native α-satellite DNA and solved its structure by the cryo-electron microscopy (cryo-EM) to 2.6 Å resolution. In comparison, the corresponding cryo-EM structure of the free CENP-A nucleosome could only reach 3.4 Å resolution. We find that scFv binds to a conserved acidic patch on the histone H2A-H2B dimer without perturbing the nucleosome structure. Our results provide an atomic resolution cryo-EM structure of a nucleosome and insight into the structure and function of the CENP-A nucleosome. The scFv approach is applicable to the structural determination of other native-like nucleosomes with distinct DNA sequences.
- Laboratory of Biochemistry and Molecular Biology, Center for Cancer Research, National Cancer Institute, National Institutes of Health, Bethesda, MD, 20892, USA.
Organizational Affiliation: