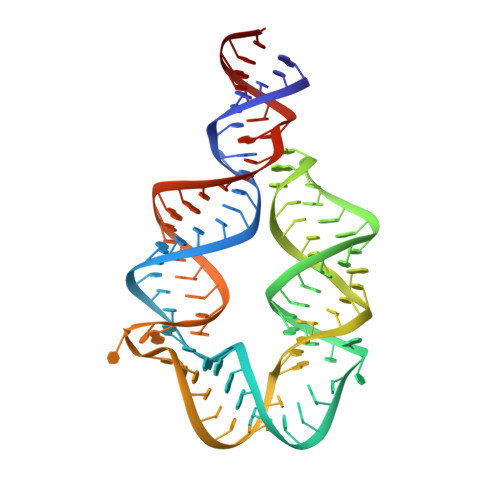
Computational design of three-dimensional RNA structure and function.
Yesselman, J.D., Eiler, D., Carlson, E.D., Gotrik, M.R., d'Aquino, A.E., Ooms, A.N., Kladwang, W., Carlson, P.D., Shi, X., Costantino, D.A., Herschlag, D., Lucks, J.B., Jewett, M.C., Kieft, J.S., Das, R.(2019) Nat Nanotechnol 14: 866-873
- PubMed: 31427748 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41565-019-0517-8
- Primary Citation Related Structures:
6DVK - PubMed Abstract:
RNA nanotechnology seeks to create nanoscale machines by repurposing natural RNA modules. The field is slowed by the current need for human intuition during three-dimensional structural design. Here, we demonstrate that three distinct problems in RNA nanotechnology can be reduced to a pathfinding problem and automatically solved through an algorithm called RNAMake. First, RNAMake discovers highly stable single-chain solutions to the classic problem of aligning a tetraloop and its sequence-distal receptor, with experimental validation from chemical mapping, gel electrophoresis, solution X-ray scattering and crystallography with 2.55 Å resolution. Second, RNAMake automatically generates structured tethers that integrate 16S and 23S ribosomal RNAs into single-chain ribosomal RNAs that remain uncleaved by ribonucleases and assemble onto messenger RNA. Third, RNAMake enables the automated stabilization of small-molecule binding RNAs, with designed tertiary contacts that improve the binding affinity of the ATP aptamer and improve the fluorescence and stability of the Spinach RNA in cell extracts and in living Escherichia coli cells.
- Department of Biochemistry, Stanford University School of Medicine, Stanford, CA, USA.
Organizational Affiliation: