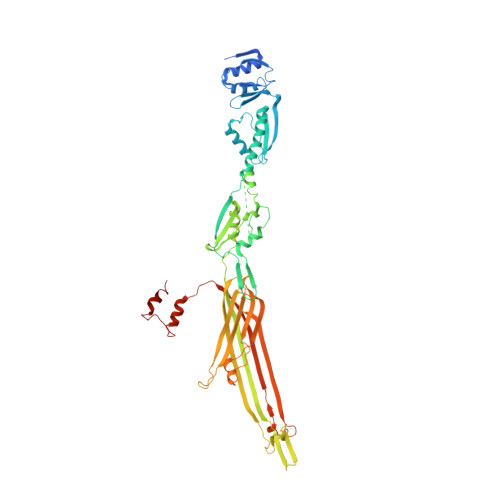
Cryo-EM analysis of the T3S injectisome reveals the structure of the needle and open secretin.
Hu, J., Worrall, L.J., Hong, C., Vuckovic, M., Atkinson, C.E., Caveney, N., Yu, Z., Strynadka, N.C.J.(2018) Nat Commun 9: 3840-3840
- PubMed: 30242280 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-06298-8
- Primary Citation Related Structures:
6DUZ, 6DV3, 6DV6, 6DWB - PubMed Abstract:
The bacterial type III secretion system, or injectisome, is a syringe shaped nanomachine essential for the virulence of many disease causing Gram-negative bacteria. At the core of the injectisome structure is the needle complex, a continuous channel formed by the highly oligomerized inner and outer membrane hollow rings and a polymerized helical needle filament which spans through and projects into the infected host cell. Here we present the near-atomic resolution structure of a needle complex from the prototypical Salmonella Typhimurium SPI-1 type III secretion system, with local masking protocols allowing for model building and refinement of the major membrane spanning components of the needle complex base in addition to an isolated needle filament. This work provides significant insight into injectisome structure and assembly and importantly captures the molecular basis for substrate induced gating in the giant outer membrane secretin portal family.
- Department of Biochemistry and Molecular Biology and the Centre for Blood Research, University of British Columbia, Vancouver, V6T 1Z3, BC, Canada.
Organizational Affiliation: