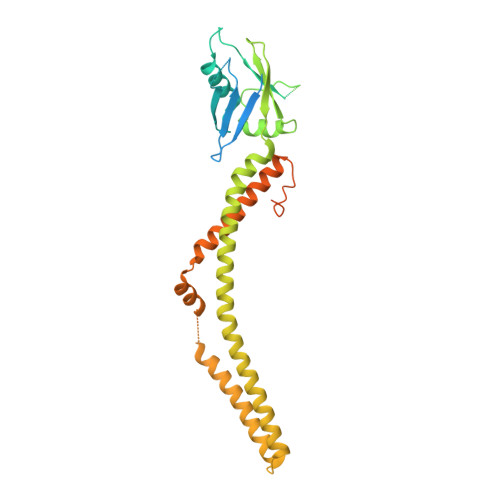
Cryo-EM structure of a mitochondrial calcium uniporter.
Yoo, J., Wu, M., Yin, Y., Herzik Jr., M.A., Lander, G.C., Lee, S.Y.(2018) Science 361: 506-511
- PubMed: 29954988 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.aar4056
- Primary Citation Related Structures:
6DT0 - PubMed Abstract:
Calcium transport plays an important role in regulating mitochondrial physiology and pathophysiology. The mitochondrial calcium uniporter (MCU) is a calcium-selective ion channel that is the primary mediator for calcium uptake into the mitochondrial matrix. Here, we present the cryo-electron microscopy structure of the full-length MCU from Neurospora crassa to an overall resolution of ~3.7 angstroms. Our structure reveals a tetrameric architecture, with the soluble and transmembrane domains adopting different symmetric arrangements within the channel. The conserved W-D-Φ-Φ-E-P-V-T-Y sequence motif of MCU pore forms a selectivity filter comprising two acidic rings separated by one helical turn along the central axis of the channel pore. The structure combined with mutagenesis gives insight into the basis of calcium recognition.
- Department of Biochemistry, Duke University School of Medicine, Durham, NC 27710, USA.
Organizational Affiliation: