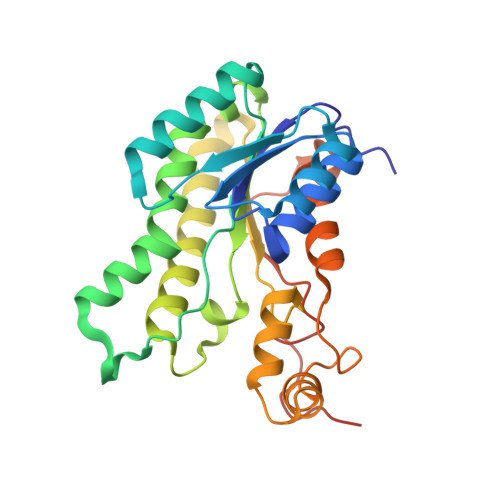
The gastrointestinal pathogen Campylobacter jejuni metabolizes sugars with potential help from commensal Bacteroides vulgatus.
Garber, J.M., Nothaft, H., Pluvinage, B., Stahl, M., Bian, X., Porfirio, S., Enriquez, A., Butcher, J., Huang, H., Glushka, J., Line, E., Gerlt, J.A., Azadi, P., Stintzi, A., Boraston, A.B., Szymanski, C.M.(2020) Commun Biol 3: 2-2
- PubMed: 31925306 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-019-0727-5
- Primary Citation Related Structures:
6DRR, 6DS1 - PubMed Abstract:
Although the gastrointestinal pathogen Campylobacter jejuni was considered asaccharolytic, >50% of sequenced isolates possess an operon for L-fucose utilization. In C. jejuni NCTC11168, this pathway confers L-fucose chemotaxis and competitive colonization advantages in the piglet diarrhea model, but the catabolic steps remain unknown. Here we solved the putative dehydrogenase structure, resembling FabG of Burkholderia multivorans. The C. jejuni enzyme, FucX, reduces L-fucose and D-arabinose in vitro and both sugars are catabolized by fuc-operon encoded enzymes. This enzyme alone confers chemotaxis to both sugars in a non-carbohydrate-utilizing C. jejuni strain. Although C. jejuni lacks fucosidases, the organism exhibits enhanced growth in vitro when co-cultured with Bacteroides vulgatus, suggesting scavenging may occur. Yet, when excess amino acids are available, C. jejuni prefers them to carbohydrates, indicating a metabolic hierarchy exists. Overall this study increases understanding of nutrient metabolism by this pathogen, and identifies interactions with other gut microbes.
- Department of Microbiology, University of Georgia, 527 Biological Sciences Bldg, Athens, GA, 30602, USA.
Organizational Affiliation: