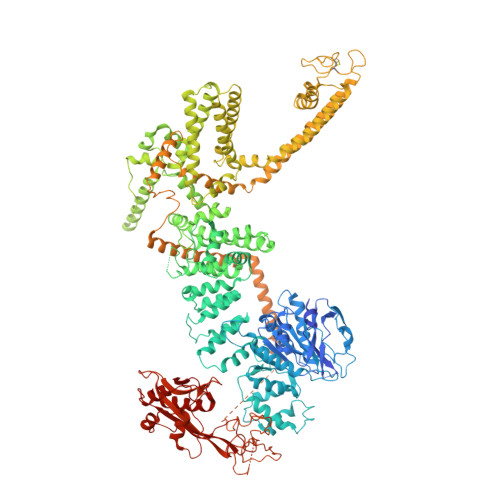
Architecture of the TRPM2 channel and its activation mechanism by ADP-ribose and calcium.
Huang, Y., Winkler, P.A., Sun, W., Lu, W., Du, J.(2018) Nature 562: 145-149
- PubMed: 30250252 Search on PubMed
- DOI: https://doi.org/10.1038/s41586-018-0558-4
- Primary Citation Related Structures:
6DRJ, 6DRK - PubMed Abstract:
Transient receptor potential melastatin 2 (TRPM2) is a calcium-permeable, non-selective cation channel that has an essential role in diverse physiological processes such as core body temperature regulation, immune response and apoptosis 1-4 . TRPM2 is polymodal and can be activated by a wide range of stimuli 1-7 , including temperature, oxidative stress and NAD + -related metabolites such as ADP-ribose (ADPR). Its activation results in both Ca 2+ entry across the plasma membrane and Ca 2+ release from lysosomes 8 , and has been linked to diseases such as ischaemia-reperfusion injury, bipolar disorder and Alzheimer's disease 9-11 . Here we report the cryo-electron microscopy structures of the zebrafish TRPM2 in the apo resting (closed) state and in the ADPR/Ca 2+ -bound active (open) state, in which the characteristic NUDT9-H domains hang underneath the MHR1/2 domain. We identify an ADPR-binding site located in the bi-lobed structure of the MHR1/2 domain. Our results provide an insight into the mechanism of activation of the TRPM channel family and define a framework for the development of therapeutic agents to treat neurodegenerative diseases and temperature-related pathological conditions.
- Van Andel Research Institute, Grand Rapids, MI, USA.
Organizational Affiliation: