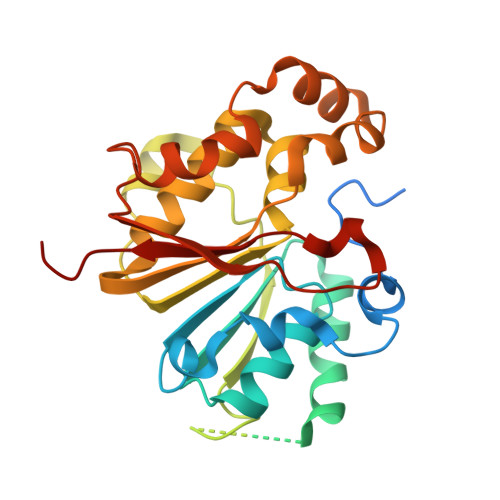
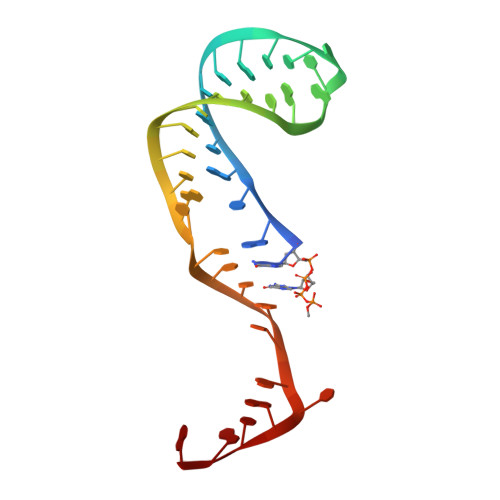
Structural basis of 7SK RNA 5'-gamma-phosphate methylation and retention by MePCE.
Yang, Y., Eichhorn, C.D., Wang, Y., Cascio, D., Feigon, J.(2019) Nat Chem Biol 15: 132-140
- PubMed: 30559425 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-018-0188-z
- Primary Citation Related Structures:
6DCB, 6DCC - PubMed Abstract:
Among RNA 5'-cap structures, γ-phosphate monomethylation is unique to a small subset of noncoding RNAs, 7SK and U6 in humans. 7SK is capped by methylphosphate capping enzyme (MePCE), which has a second nonenzymatic role as a core component of the 7SK ribonuclear protein (RNP), an essential regulator of RNA transcription. We report 2.0- and 2.1-Å X-ray crystal structures of the human MePCE methyltransferase domain bound to S-adenosylhomocysteine (SAH) and uncapped or capped 7SK substrates, respectively. 7SK recognition is achieved by protein contacts to a 5'-hairpin-single-stranded RNA region, thus explaining MePCE's specificity for 7SK and U6. The structures reveal SAH and product RNA in a near-transition-state geometry. Unexpectedly, binding experiments showed that MePCE has higher affinity for capped versus uncapped 7SK, and kinetic data support a model of slow product release. This work reveals the molecular mechanism of methyl transfer and 7SK retention by MePCE for subsequent assembly of 7SK RNP.
- Department of Chemistry and Biochemistry, University of California, Los Angeles, Los Angeles, CA, USA.
Organizational Affiliation: