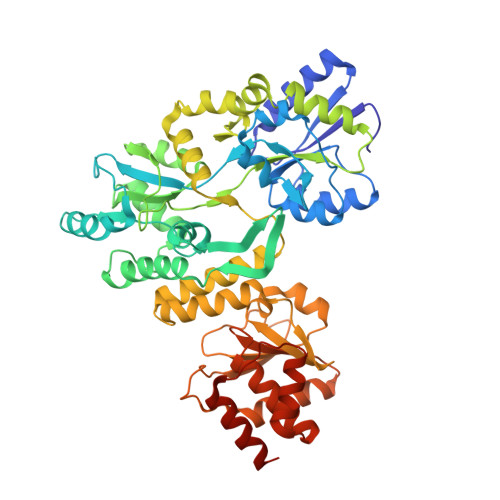
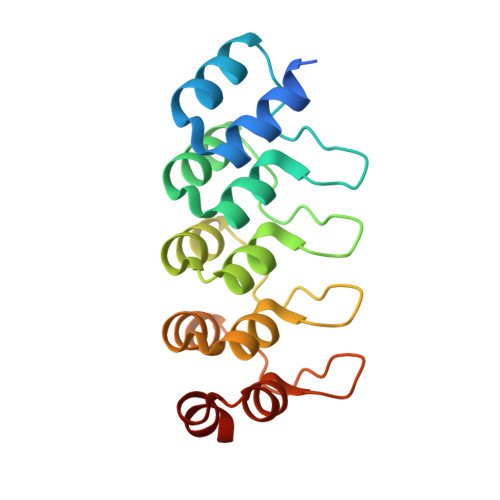
MBP-binding DARPins facilitate the crystallization of an MBP fusion protein.
Gumpena, R., Lountos, G.T., Waugh, D.S.(2018) Acta Crystallogr F Struct Biol Commun 74: 549-557
- PubMed: 30198887 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X18009901
- Primary Citation Related Structures:
6D65, 6D66, 6D67 - PubMed Abstract:
The production of high-quality crystals is the main bottleneck in determining the structures of proteins using X-ray crystallography. In addition to being recognized as a very effective solubility-enhancing fusion partner, Escherichia coli maltose-binding protein (MBP) has also been successfully employed as a `fixed-arm' crystallization chaperone in more than 100 cases. Here, it is reported that designed ankyrin-repeat proteins (DARPins) that bind with high affinity to MBP can promote the crystallization of an MBP fusion protein when the fusion protein alone fails to produce diffraction-quality crystals. As a proof of principle, three different co-crystal structures of MBP fused to the catalytic domain of human dual-specificity phosphatase 1 in complex with DARPins are reported.
- Macromolecular Crystallography Laboratory, Center for Cancer Research, National Cancer Institute at Frederick, Frederick, MD 21702, USA.
Organizational Affiliation: