A subset of HLA-I peptides are not genomically templated: Evidence for cis- and trans-spliced peptide ligands.
Faridi, P., Li, C., Ramarathinam, S.H., Vivian, J.P., Illing, P.T., Mifsud, N.A., Ayala, R., Song, J., Gearing, L.J., Hertzog, P.J., Ternette, N., Rossjohn, J., Croft, N.P., Purcell, A.W.(2018) Sci Immunol 3
- PubMed: 30315122 Search on PubMed
- DOI: https://doi.org/10.1126/sciimmunol.aar3947
- Primary Citation Related Structures:
6D29, 6D2B, 6D2R, 6D2T - PubMed Abstract:
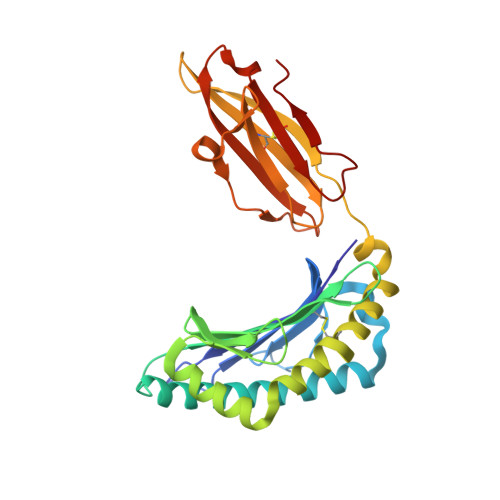
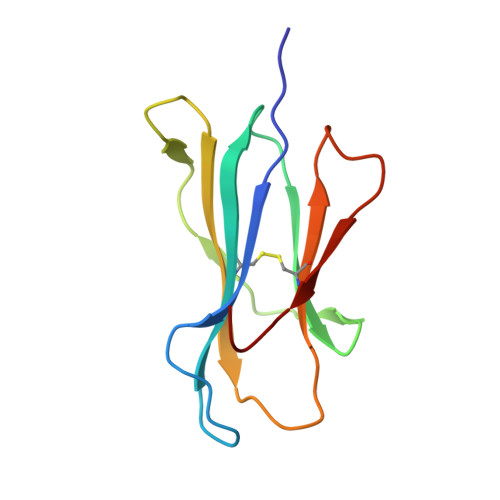
The diversity of peptides displayed by class I human leukocyte antigen (HLA) plays an essential role in T cell immunity. The peptide repertoire is extended by various posttranslational modifications, including proteasomal splicing of peptide fragments from distinct regions of an antigen to form nongenomically templated cis-spliced sequences. Previously, it has been suggested that a fraction of the immunopeptidome constitutes such cis-spliced peptides; however, because of computational limitations, it has not been possible to assess whether trans-spliced peptides (i.e., the fusion of peptide segments from distinct antigens) are also bound and presented by HLA molecules, and if so, in what proportion. Here, we have developed and applied a bioinformatic workflow and demonstrated that trans-spliced peptides are presented by HLA-I, and their abundance challenges current models of proteasomal splicing that predict cis-splicing as the most probable outcome. These trans-spliced peptides display canonical HLA-binding sequence features and are as frequently identified as cis-spliced peptides found bound to a number of different HLA-A and HLA-B allotypes. Structural analysis reveals that the junction between spliced peptides is highly solvent exposed and likely to participate in T cell receptor interactions. These results highlight the unanticipated diversity of the immunopeptidome and have important implications for autoimmunity, vaccine design, and immunotherapy.
- Infection and Immunity Program and Department of Biochemistry and Molecular Biology, Biomedicine Discovery Institute, Monash University, Clayton, Victoria 3800, Australia.
Organizational Affiliation: