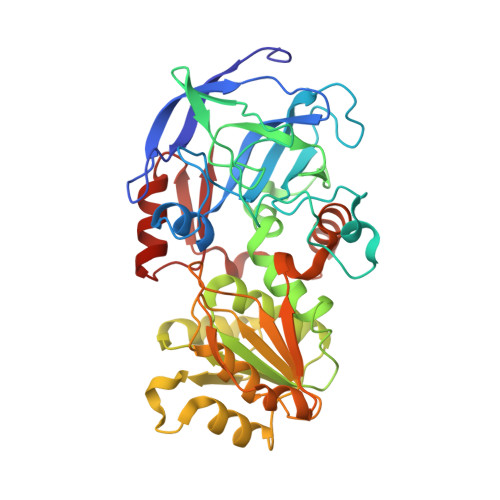
Substitutions of a buried glutamate residue hinder the conformational change in horse liver alcohol dehydrogenase and yield a surprising complex with endogenous 3'-Dephosphocoenzyme A.
Kim, Y.H., Gogerty, D.S., Plapp, B.V.(2018) Arch Biochem Biophys 653: 97-106
- PubMed: 30018019 Search on PubMed
- DOI: https://doi.org/10.1016/j.abb.2018.07.003
- Primary Citation Related Structures:
6CXX, 6CY3 - PubMed Abstract:
Glu-267 is highly conserved in alcohol dehydrogenases and buried as a negatively-charged residue in a loop of the NAD coenzyme binding domain. Glu-267 might have a structural role and contribute to a rate-promoting vibration that facilitates catalysis. Substitutions of Glu-267 with histidine or asparagine residues increase the dissociation constants for the coenzymes (NAD + by ∼40-fold, NADH by ∼200-fold) and significantly decrease catalytic efficiencies by 16-1200-fold various substrates and substituted enzymes. The turnover numbers modestly change with the substitutions, but hydride transfer is at least partially rate-limiting for turnover for alcohol oxidation. X-ray structures of the E267H and E267 N enzymes are similar to the apoenzyme (open) conformation of the wild-type enzyme, and the substitutions are accommodated by local changes in the structure. Surprisingly, the E267H and E267 N enzymes have endogenous (from the expression in E. coli) 3'-dephosphocoenzyme A bound in the active site with the ADP moiety in the NAD binding site and the pantethiene sulfhydryl bound to the catalytic zinc. The kinetics and crystallography show that the substitutions of Glu-267 hinder the conformational change, which occurs when wild-type enzyme binds coenzymes, and affect productive binding of substrates.
- Department of Biochemistry, The University of Iowa, Iowa City, IA, 52242, USA. Electronic address: ykim@uwa.edu.
Organizational Affiliation: