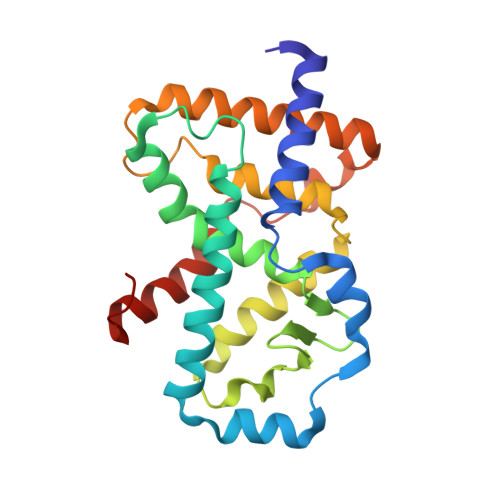
Identification and biological evaluation of thiazole-based inverse agonists of ROR gamma t.
Gege, C., Cummings, M.D., Albers, M., Kinzel, O., Kleymann, G., Schluter, T., Steeneck, C., Nelen, M.I., Milligan, C., Spurlino, J., Xue, X., Leonard, K., Edwards, J.P., Fourie, A., Goldberg, S.D., Hoffmann, T.(2018) Bioorg Med Chem Lett 28: 1446-1455
- PubMed: 29631962 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2018.03.093
- Primary Citation Related Structures:
6CVH - PubMed Abstract:
The nuclear receptor retinoic acid receptor-related orphan receptor gamma t (RORγt) is a transcription factor that drives Th17 cell differentiation and IL-17 production in both innate and adaptive immune cells. The IL-23/IL-17 pathway is implicated in major autoimmune and inflammatory diseases. RORγt lies at the core of this pathway and represents an attractive opportunity for intervention with a small molecule. Despite diverse chemical series having been reported, combining high potency and nuclear receptor selectivity with good physicochemical properties remains a challenging endeavor in the field of RORγt drug discovery. We describe the discovery and evaluation of a new class of potent and selective RORγt inverse agonists based on a thiazole core. Acid analog 1j demonstrated oral bioavailability in rats and was potent in a human whole blood assay, suggesting potential utility in treating autoimmune and inflammatory diseases such as psoriasis. X-ray crystallographic data helped to elucidate the molecular mechanism for RORγt inhibition with this series.
- Phenex Pharmaceuticals AG, Waldhofer Strasse 104, 69123 Heidelberg, Germany. Electronic address: christian.gege@web.de.
Organizational Affiliation: