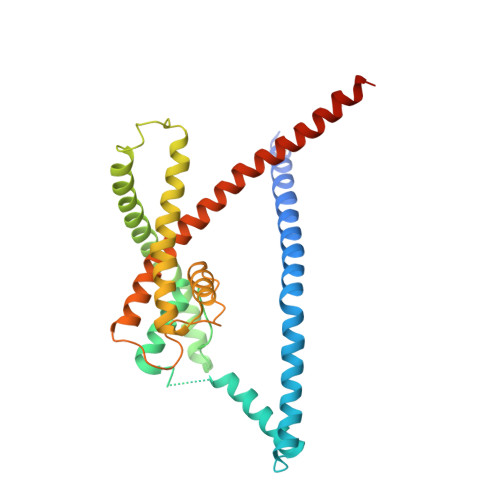
K2P2.1 (TREK-1)-activator complexes reveal a cryptic selectivity filter binding site.
Lolicato, M., Arrigoni, C., Mori, T., Sekioka, Y., Bryant, C., Clark, K.A., Minor, D.L.(2017) Nature 547: 364-368
- PubMed: 28693035 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature22988
- Primary Citation Related Structures:
6CQ6, 6CQ8, 6CQ9 - PubMed Abstract:
Polymodal thermo- and mechanosensitive two-pore domain potassium (K 2P ) channels of the TREK subfamily generate 'leak' currents that regulate neuronal excitability, respond to lipids, temperature and mechanical stretch, and influence pain, temperature perception and anaesthetic responses. These dimeric voltage-gated ion channel (VGIC) superfamily members have a unique topology comprising two pore-forming regions per subunit. In contrast to other potassium channels, K 2P channels use a selectivity filter 'C-type' gate as the principal gating site. Despite recent advances, poor pharmacological profiles of K 2P channels limit mechanistic and biological studies. Here we describe a class of small-molecule TREK activators that directly stimulate the C-type gate by acting as molecular wedges that restrict interdomain interface movement behind the selectivity filter. Structures of K 2P 2.1 (also known as TREK-1) alone and with two selective K 2P 2.1 (TREK-1) and K 2P 10.1 (TREK-2) activators-an N-aryl-sulfonamide, ML335, and a thiophene-carboxamide, ML402-define a cryptic binding pocket unlike other ion channel small-molecule binding sites and, together with functional studies, identify a cation-π interaction that controls selectivity. Together, our data reveal a druggable K 2P site that stabilizes the C-type gate 'leak mode' and provide direct evidence for K 2P selectivity filter gating.
- Cardiovascular Research Institute, University of California, San Francisco, California 941158-9001, USA.
Organizational Affiliation: