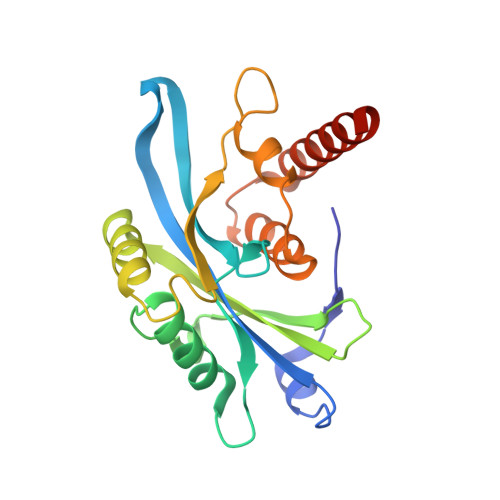
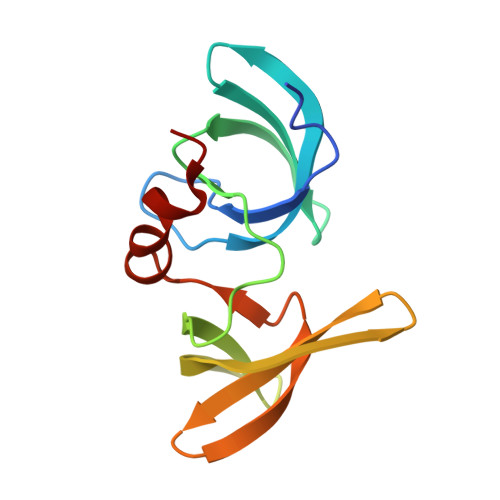
Mechanism of 53BP1 activity regulation by RNA-binding TIRR and a designer protein.
Botuyan, M.V., Cui, G., Drane, P., Oliveira, C., Detappe, A., Brault, M.E., Parnandi, N., Chaubey, S., Thompson, J.R., Bragantini, B., Zhao, D., Chapman, J.R., Chowdhury, D., Mer, G.(2018) Nat Struct Mol Biol 25: 591-600
- PubMed: 29967538 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-018-0083-z
- Primary Citation Related Structures:
6CO1, 6CO2, 6D0L - PubMed Abstract:
Dynamic protein interaction networks such as DNA double-strand break (DSB) signaling are modulated by post-translational modifications. The DNA repair factor 53BP1 is a rare example of a protein whose post-translational modification-binding function can be switched on and off. 53BP1 is recruited to DSBs by recognizing histone lysine methylation within chromatin, an activity directly inhibited by the 53BP1-binding protein TIRR. X-ray crystal structures of TIRR and a designer protein bound to 53BP1 now reveal a unique regulatory mechanism in which an intricate binding area centered on an essential TIRR arginine residue blocks the methylated-chromatin-binding surface of 53BP1. A 53BP1 separation-of-function mutation that abolishes TIRR-mediated regulation in cells renders 53BP1 hyperactive in response to DSBs, highlighting the key inhibitory function of TIRR. This 53BP1 inhibition is relieved by TIRR-interacting RNA molecules, providing proof-of-principle of RNA-triggered 53BP1 recruitment to DSBs.
- Department of Biochemistry and Molecular Biology, Mayo Clinic, Rochester, MN, USA.
Organizational Affiliation: