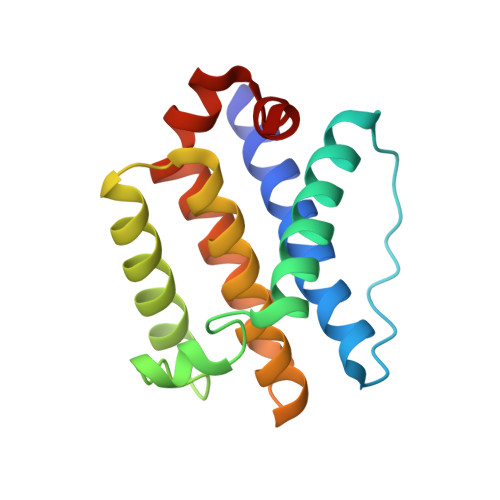
Intrinsic Instability of BOK Enables Membrane Permeabilization in Apoptosis.
Zheng, J.H., Grace, C.R., Guibao, C.D., McNamara, D.E., Llambi, F., Wang, Y.M., Chen, T., Moldoveanu, T.(2018) Cell Rep 23: 2083-2094.e6
- PubMed: 29768206 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2018.04.060
- Primary Citation Related Structures:
6CKV - PubMed Abstract:
The effector B cell lymphoma-2 (BCL-2) protein BCL-2 ovarian killer (BOK) induces mitochondrial outer membrane permeabilization (MOMP) to initiate apoptosis upon inhibition of the proteasome. How BOK mediates MOMP is mechanistically unknown. The NMR structure of the BCL-2 core of human BOK reveals a conserved architecture with an atypical hydrophobic groove that undergoes conformational exchange. Remarkably, the BCL-2 core of BOK spontaneously associates with purified mitochondria to release cytochrome c in MOMP assays. Alanine substitution of a unique glycine in helix α1 stabilizes BOK, as shown by thermal shift and urea denaturation analyses, and significantly inhibits MOMP, liposome permeabilization, and cell death. Activated BID does not activate WT BOK or the stabilized alanine mutant to promote cell death. We propose that BOK-mediated membrane permeabilization is governed in part by its unique metastability of the hydrophobic groove and helix α1 and not through activation by BH3 ligands.
- Department of Structural Biology, St. Jude Children's Research Hospital, Memphis, TN 38105, USA; Department of Chemical Biology and Therapeutics, St. Jude Children's Research Hospital, Memphis, TN 38105, USA.
Organizational Affiliation: