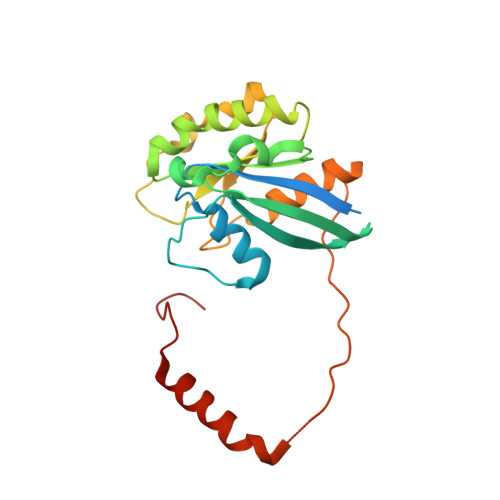
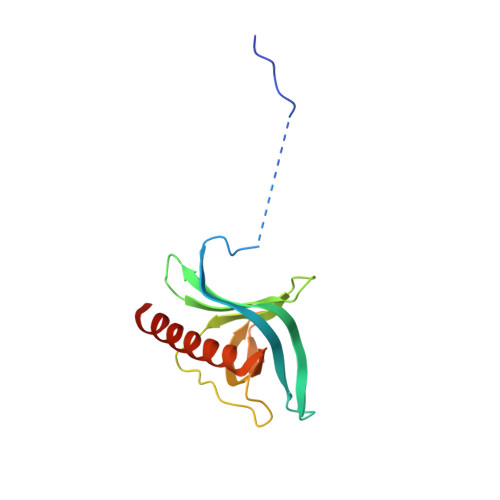
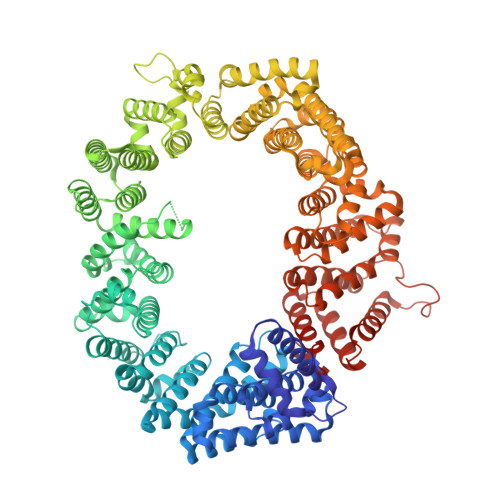
Correlation of CRM1-NES affinity with nuclear export activity.
Fu, S.C., Fung, H.Y.J., Cagatay, T., Baumhardt, J., Chook, Y.M.(2018) Mol Biol Cell 29: 2037-2044
- PubMed: 29927350 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1091/mbc.E18-02-0096
- Primary Citation Related Structures:
6CIT - PubMed Abstract:
CRM1 (Exportin1/XPO1) exports hundreds of broadly functioning protein cargoes out of the cell nucleus by binding to their classical nuclear export signals (NESs). The 8- to 15-amino-acid-long NESs contain four to five hydrophobic residues and are highly diverse in both sequence and CRM1-bound structure. Here we examine the relationship between nuclear export activities of 24 different NES peptides in cells and their CRM1-NES affinities. We found that binding affinity and nuclear export activity are linearly correlated for NESs with dissociation constants ( K d s) between tens of nanomolar to tens of micromolar. NESs with K d s outside this range have significantly reduced nuclear export activities. These include two unusually tight-binding peptides, one from the nonstructural protein 2 of murine minute virus (MVM NS2) and the other a mutant of the protein kinase A inhibitor (PKI) NES. The crystal structure of CRM1-bound MVM NS2 NES suggests that extraordinarily tight CRM1 binding arises from intramolecular contacts within the NES that likely stabilizes the CRM1-bound conformation in free peptides. This mechanistic understanding led to the design of two novel peptide inhibitors that bind CRM1 with picomolar affinity.
- Department of Pharmacology, University of Texas Southwestern Medical Center, Dallas, TX 75390.
Organizational Affiliation: