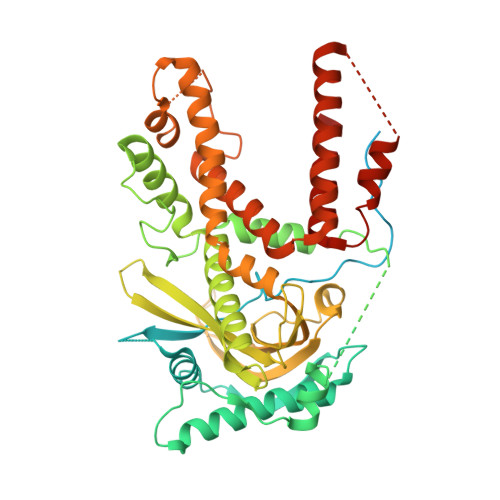
CRYSTAL STRUCTURE OF POLYMERASE ACID PROTEIN (PA) FROM INFLUENZA A VIRUS, WILSON-SMITH/1933 (H1N1) BOUND TO FRAGMENT HIT BSI-70565 1-{1-[4-FLUOROPHENYL)METHYL]-2-METHYL-1H-IMIDAZOL-4-YL}ETHAN-1-ONE
Seattle Structural Genomics Center for Infectious Disease (SSGCID), Davies, D.R., Abendroth, J., Lorimer, D., Horanyi, P.S., Edwards, T.E.To be published.