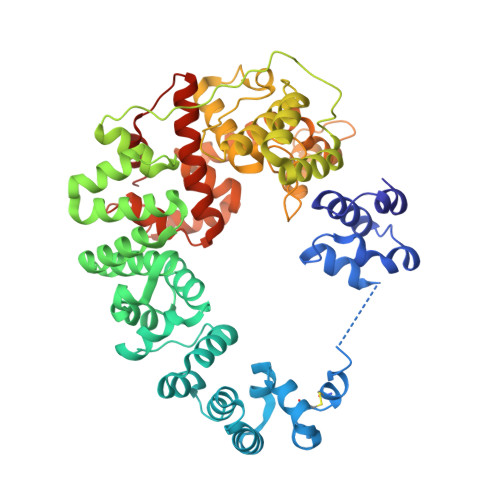
Structural studies and molecular dynamics simulations suggest a processive mechanism of exolytic lytic transglycosylase from Campylobacter jejuni.
Vijayaraghavan, J., Kumar, V., Krishnan, N.P., Kaufhold, R.T., Zeng, X., Lin, J., van den Akker, F.(2018) PLoS One 13: e0197136-e0197136
- PubMed: 29758058 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0197136
- Primary Citation Related Structures:
6CF8, 6CF9, 6CFC - PubMed Abstract:
The bacterial soluble lytic transglycosylase (LT) breaks down the peptidoglycan (PG) layer during processes such as cell division. We present here crystal structures of the soluble LT Cj0843 from Campylobacter jejuni with and without bulgecin A inhibitor in the active site. Cj0843 has a doughnut shape similar but not identical to that of E. coli SLT70. The C-terminal catalytic domain is preceded by an L-domain, a large helical U-domain, a flexible linker, and a small N-terminal NU-domain. The flexible linker allows the NU-domain to reach over and complete the circular shape, using residues conserved in the Epsilonproteobacteria LT family. The inner surface of the Cj0843 doughnut is mostly positively charged including a pocket that has 8 Arg/Lys residues. Molecular dynamics simulations with PG strands revealed a potential functional role for this pocket in anchoring the negatively charged terminal tetrapeptide of the PG during several steps in the reaction including homing and aligning the PG strand for exolytic cleavage, and subsequent ratcheting of the PG strand to enhance processivity in degrading PG strands.
- Department of Biochemistry, Case Western Reserve University, Cleveland, OH, United States of America.
Organizational Affiliation: