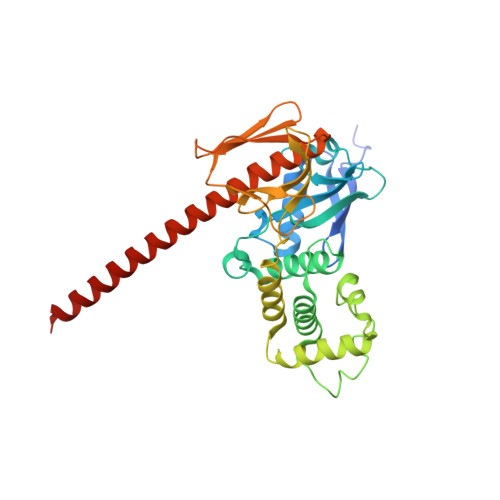
Lipid binding promotes the open conformation and tumor-suppressive activity of neurofibromin 2.
Chinthalapudi, K., Mandati, V., Zheng, J., Sharff, A.J., Bricogne, G., Griffin, P.R., Kissil, J., Izard, T.(2018) Nat Commun 9: 1338-1338
- PubMed: 29626191 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-03648-4
- Primary Citation Related Structures:
6CDS - PubMed Abstract:
Neurofibromatosis type 2 (NF2) is a tumor-forming disease of the nervous system caused by deletion or by loss-of-function mutations in NF2, encoding the tumor suppressing protein neurofibromin 2 (also known as schwannomin or merlin). Neurofibromin 2 is a member of the ezrin, radixin, moesin (ERM) family of proteins regulating the cytoskeleton and cell signaling. The correlation of the tumor-suppressive function and conformation (open or closed) of neurofibromin 2 has been subject to much speculation, often based on extrapolation from other ERM proteins, and controversy. Here we show that lipid binding results in the open conformation of neurofibromin 2 and that lipid binding is necessary for inhibiting cell proliferation. Collectively, our results provide a mechanism in which the open conformation is unambiguously correlated with lipid binding and localization to the membrane, which are critical for the tumor-suppressive function of neurofibromin 2, thus finally reconciling the long-standing conformation and function debate.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, Jupiter, FL, 33458, USA.
Organizational Affiliation: