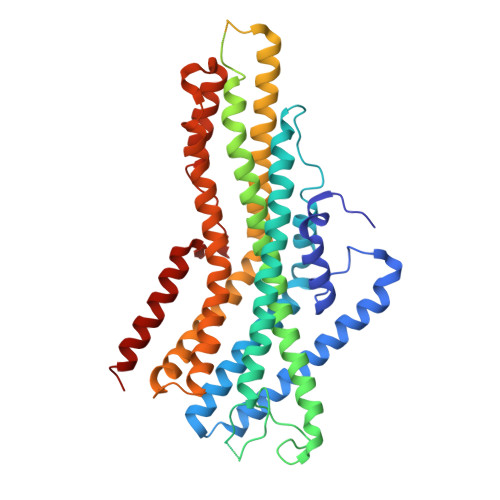
Cryo-EM structure of the insect olfactory receptor Orco.
Butterwick, J.A., Del Marmol, J., Kim, K.H., Kahlson, M.A., Rogow, J.A., Walz, T., Ruta, V.(2018) Nature 560: 447-452
- PubMed: 30111839 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-018-0420-8
- Primary Citation Related Structures:
6C70 - PubMed Abstract:
The olfactory system must recognize and discriminate amongst an enormous variety of chemicals in the environment. To contend with such diversity, insects have evolved a family of odorant-gated ion channels comprised of a highly conserved co-receptor (Orco) and a divergent odorant receptor (OR) that confers chemical specificity. Here, we present the single-particle cryo-electron microscopy structure of an Orco homomer from the parasitic fig wasp Apocrypta bakeri at 3.5 Å resolution, providing structural insight into this receptor family. Orco possesses a novel channel architecture, with four subunits symmetrically arranged around a central pore that diverges into four lateral conduits that open to the cytosol. The Orco tetramer has few inter-subunit interactions within the membrane and is bound together by a small cytoplasmic anchor domain. The minimal sequence conservation among ORs maps largely to the pore and anchor domain, shedding light on how the architecture of this receptor family accommodates its remarkable sequence diversity and facilitates the evolution of odour tuning.
- Laboratory of Neurophysiology and Behavior, The Rockefeller University, New York, NY, USA.
Organizational Affiliation: