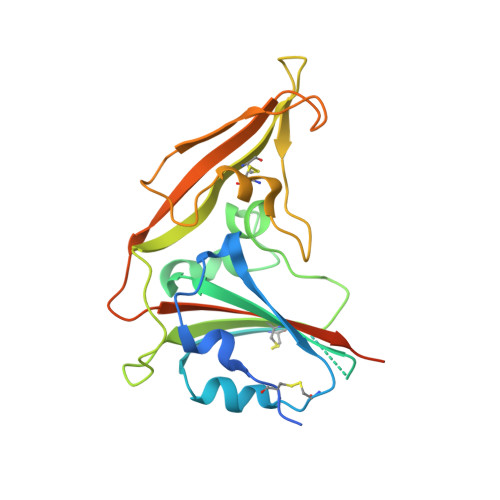
Importance of Neutralizing Monoclonal Antibodies Targeting Multiple Antigenic Sites on the Middle East Respiratory Syndrome Coronavirus Spike Glycoprotein To Avoid Neutralization Escape.
Wang, L., Shi, W., Chappell, J.D., Joyce, M.G., Zhang, Y., Kanekiyo, M., Becker, M.M., van Doremalen, N., Fischer, R., Wang, N., Corbett, K.S., Choe, M., Mason, R.D., Van Galen, J.G., Zhou, T., Saunders, K.O., Tatti, K.M., Haynes, L.M., Kwong, P.D., Modjarrad, K., Kong, W.P., McLellan, J.S., Denison, M.R., Munster, V.J., Mascola, J.R., Graham, B.S.(2018) J Virol 92
- PubMed: 29514901 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.02002-17
- Primary Citation Related Structures:
6C6X, 6C6Y, 6C6Z - PubMed Abstract:
Middle East respiratory syndrome coronavirus (MERS-CoV) causes a highly lethal pulmonary infection with ∼35% mortality. The potential for a future pandemic originating from animal reservoirs or health care-associated events is a major public health concern. There are no vaccines or therapeutic agents currently available for MERS-CoV. Using a probe-based single B cell cloning strategy, we have identified and characterized multiple neutralizing monoclonal antibodies (MAbs) specifically binding to the receptor-binding domain (RBD) or S1 (non-RBD) regions from a convalescent MERS-CoV-infected patient and from immunized rhesus macaques. RBD-specific MAbs tended to have greater neutralizing potency than non-RBD S1-specific MAbs. Six RBD-specific and five S1-specific MAbs could be sorted into four RBD and three non-RBD distinct binding patterns, based on competition assays, mapping neutralization escape variants, and structural analysis. We determined cocrystal structures for two MAbs targeting the RBD from different angles and show they can bind the RBD only in the "out" position. We then showed that selected RBD-specific, non-RBD S1-specific, and S2-specific MAbs given prophylactically prevented MERS-CoV replication in lungs and protected mice from lethal challenge. Importantly, combining RBD- and non-RBD MAbs delayed the emergence of escape mutations in a cell-based virus escape assay. These studies identify MAbs targeting different antigenic sites on S that will be useful for defining mechanisms of MERS-CoV neutralization and for developing more effective interventions to prevent or treat MERS-CoV infections. IMPORTANCE MERS-CoV causes a highly lethal respiratory infection for which no vaccines or antiviral therapeutic options are currently available. Based on continuing exposure from established reservoirs in dromedary camels and bats, transmission of MERS-CoV into humans and future outbreaks are expected. Using structurally defined probes for the MERS-CoV spike glycoprotein (S), the target for neutralizing antibodies, single B cells were sorted from a convalescent human and immunized nonhuman primates (NHPs). MAbs produced from paired immunoglobulin gene sequences were mapped to multiple epitopes within and outside the receptor-binding domain (RBD) and protected against lethal MERS infection in a murine model following passive immunization. Importantly, combining MAbs targeting distinct epitopes prevented viral neutralization escape from RBD-directed MAbs. These data suggest that antibody responses to multiple domains on CoV spike protein may improve immunity and will guide future vaccine and therapeutic development efforts.
- Vaccine Research Center, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, Maryland, USA.
Organizational Affiliation: