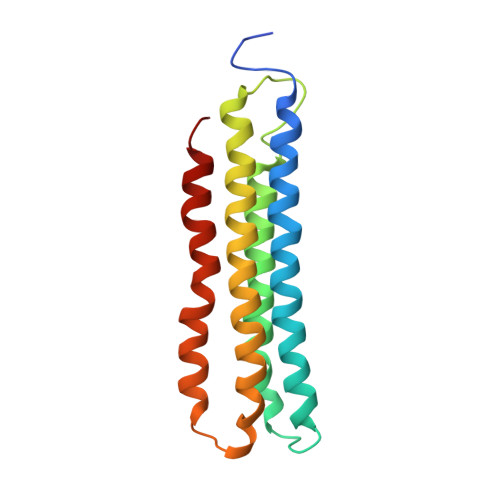
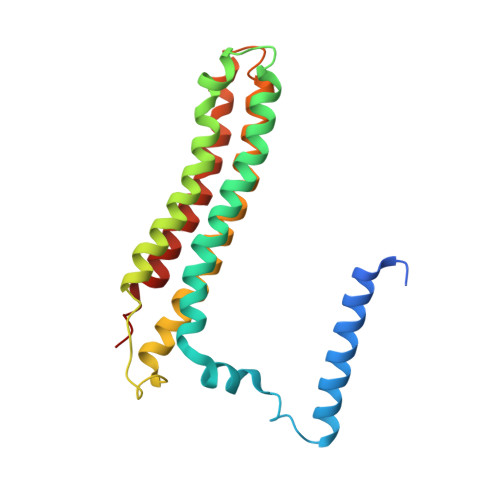
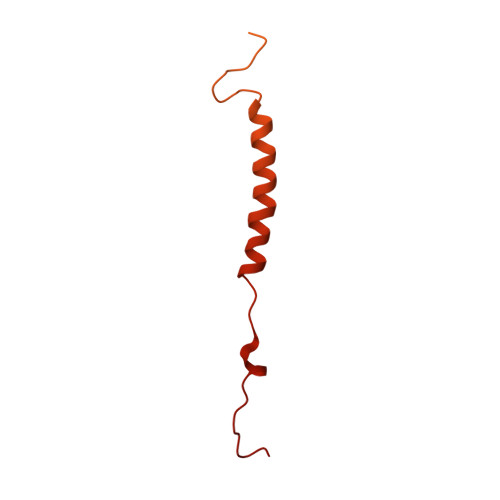
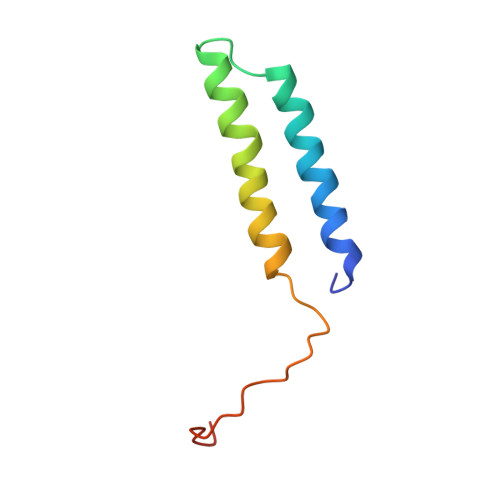
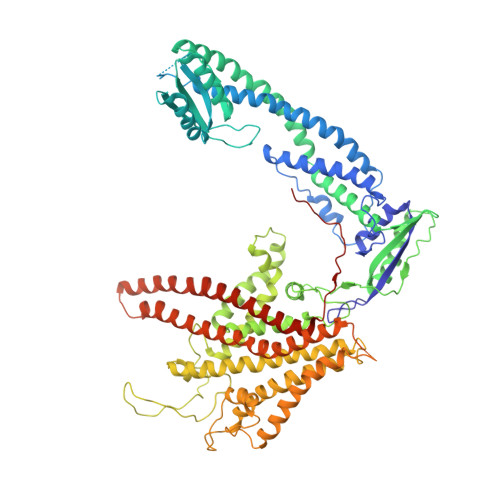
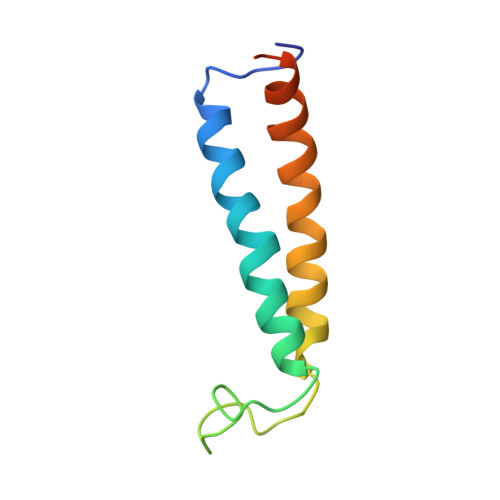
The 3.5- angstrom CryoEM Structure of Nanodisc-Reconstituted Yeast Vacuolar ATPase VoProton Channel.
Roh, S.H., Stam, N.J., Hryc, C.F., Couoh-Cardel, S., Pintilie, G., Chiu, W., Wilkens, S.(2018) Mol Cell 69: 993-1004.e3
- PubMed: 29526695 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2018.02.006
- Primary Citation Related Structures:
6C6L - PubMed Abstract:
The molecular mechanism of transmembrane proton translocation in rotary motor ATPases is not fully understood. Here, we report the 3.5-Å resolution cryoEM structure of the lipid nanodisc-reconstituted V o proton channel of the yeast vacuolar H + -ATPase, captured in a physiologically relevant, autoinhibited state. The resulting atomic model provides structural detail for the amino acids that constitute the proton pathway at the interface of the proteolipid ring and subunit a. Based on the structure and previous mutagenesis studies, we propose the chemical basis of transmembrane proton transport. Moreover, we discovered that the C terminus of the assembly factor Voa1 is an integral component of mature V o . Voa1's C-terminal transmembrane α helix is bound inside the proteolipid ring, where it contributes to the stability of the complex. Our structure rationalizes possible mechanisms by which mutations in human V o can result in disease phenotypes and may thus provide new avenues for therapeutic interventions.
- Department of Bioengineering and James H. Clark Center, Stanford University, Stanford, CA 94305, USA; Biosciences Division, SLAC National Accelerator Laboratory, Menlo Park, CA 94025, USA.
Organizational Affiliation: