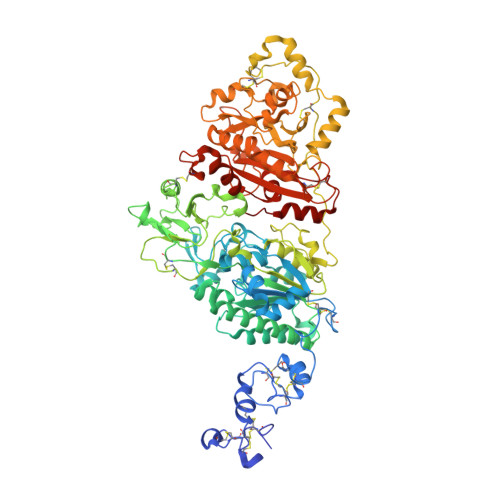
Structural basis for nucleotide recognition by the ectoenzyme CD203c.
Gorelik, A., Randriamihaja, A., Illes, K., Nagar, B.(2018) FEBS J 285: 2481-2494
- PubMed: 29717535 Search on PubMed
- DOI: https://doi.org/10.1111/febs.14489
- Primary Citation Related Structures:
6C01, 6C02 - PubMed Abstract:
The ecto-nucleotide pyrophosphatase/phosphodiesterase (NPP) enzyme family modulates purinergic signaling by degrading extracellular nucleotides. CD203c (NPP3, ENPP3) regulates the inflammatory response of basophils via ATP hydrolysis and is a marker for allergen sensitivity on the surface of these cells. Multiple other roles and substrates have also been proposed for this protein. In order to gain insight into its molecular functions, we determined the crystal structure of human NPP3 as well as its complex with an ATP analog. The enzyme exhibits little preference for nucleobase type, and forms specific contacts with the alpha and beta phosphate groups of its ligands. Dimerization of the protein does not affect its catalytic activity. These findings expand our understanding of substrate recognition within the NPP family. Structural data are available in the Protein Data Bank under the accession numbers 6C01 (human NPP3) and 6C02 (human NPP3 T205A N594S with AMPCPP).
- Department of Biochemistry and Groupe de Recherche Axé sur la Structure des Protéines, McGill University, Montreal, Quebec, Canada.
Organizational Affiliation: