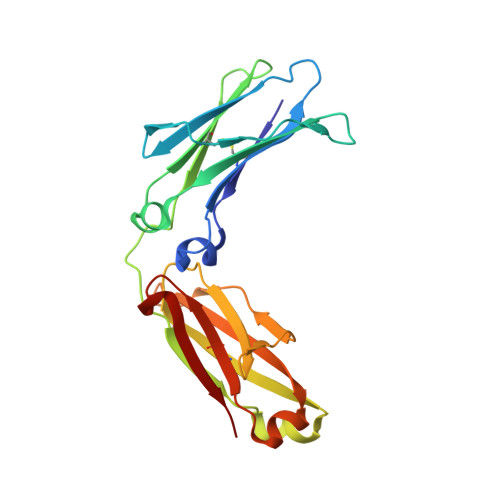
Global conformational changes in IgG-Fc upon mutation of the FcRn-binding site are not associated with altered antibody-dependent effector functions.
Burvenich, I.J.G., Farrugia, W., Liu, Z., Makris, D., King, D., Gloria, B., Perani, A., Allan, L.C., Scott, A.M., Ramsland, P.A.(2018) Biochem J 475: 2179-2190
- PubMed: 29794155 Search on PubMed
- DOI: https://doi.org/10.1042/BCJ20180139
- Primary Citation Related Structures:
6BZ4 - PubMed Abstract:
Antibody engineering is important for many diagnostic and clinical applications of monoclonal antibodies. We recently reported a series of fragment crystallizable (Fc) mutations targeting the neonatal Fc receptor (FcRn) site on a Lewis Y (Le y ) binding IgG1, hu3S193. The hu3S193 variants displayed shortened in vivo half-lives and may have potential for radioimaging or radiotherapy of Le y -positive tumors. Here, we report Fc crystal structures of wild-type hu3S193, seven FcRn-binding site variants, and a variant lacking C1q binding or complement-dependent cytotoxicity (CDC) activity. The Fc conformation of the FcRn-binding sites was similar for wild-type and all mutants of hu3S193 Fc, which suggests that FcRn interactions were directly affected by the amino acid substitutions. The C1q-binding site mutant Fc was nearly identical with the wild-type Fc. Surprisingly, several hu3S193 Fc variants showed large changes in global structure compared with wild-type Fc. All hu3S193 Fc mutants had similar antibody-dependent cellular cytotoxicity, despite some with conformations expected to diminish Fc gamma receptor binding. Several hu3S193 variants displayed altered CDC, but there was no correlation with the different Fc conformations. All versions of hu3S193, except the C1q-binding site mutant, bound C1q, suggesting that the altered CDC of some variants could result from different propensities to form IgG hexamers after engaging Le y on target cells. Overall, our findings support the concept that the antibody Fc is both flexible and mobile in solution. Structure-based design approaches should take into account the conformational plasticity of the Fc when engineering antibodies with optimal effector properties.
- Tumour Targeting Laboratory, Olivia Newton-John Cancer Research Institute, Melbourne, Australia.
Organizational Affiliation: