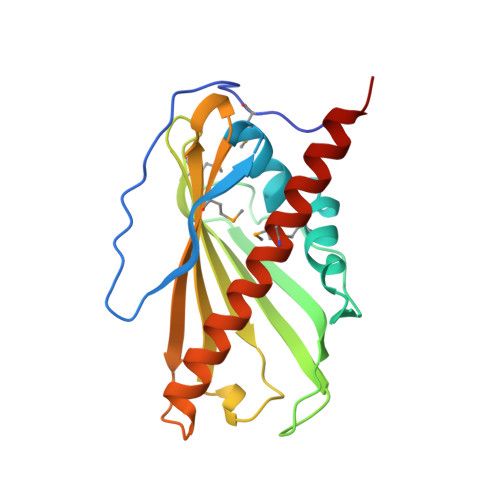
Structural basis of sterol binding and transport by a yeast StARkin domain.
Jentsch, J.A., Kiburu, I., Pandey, K., Timme, M., Ramlall, T., Levkau, B., Wu, J., Eliezer, D., Boudker, O., Menon, A.K.(2018) J Biological Chem 293: 5522-5531
- PubMed: 29463678 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA118.001881
- Primary Citation Related Structures:
6BYD, 6BYM - PubMed Abstract:
The StARkin superfamily comprises proteins with steroidogenic acute regulatory protein-related lipid transfer (StART) domains that are implicated in intracellular, non-vesicular lipid transport. A new family of membrane-anchored StARkins was recently identified, including six members, Lam1-Lam6, in the yeast Saccharomyces cerevisiae. Lam1-Lam4 are anchored to the endoplasmic reticulum (ER) membrane at sites where the ER is tethered to the plasma membrane and proposed to be involved in sterol homeostasis in yeast. To better understand the biological roles of these proteins, we carried out a structure-function analysis of the second StARkin domain of Lam4, here termed Lam4S2. NMR experiments indicated that Lam4S2 undergoes specific conformational changes upon binding sterol, and fluorescence-based assays revealed that it catalyzes sterol transport between vesicle populations in vitro , exhibiting a preference for vesicles containing anionic lipids. Using such vesicles, we found that sterols are transported at a rate of ∼50 molecules per Lam4S2 per minute. Crystal structures of Lam4S2, with and without bound sterol, revealed a largely hydrophobic but surprisingly accessible sterol-binding pocket with the 3-OH group of the sterol oriented toward its base. Single or multiple alanine or aspartic acid replacements of conserved lysine residues in a basic patch on the surface of Lam4S2 near the likely sterol entry/egress site strongly attenuated sterol transport. Our results suggest that Lam4S2 engages anionic membranes via a basic surface patch, enabling "head-first" entry of sterol into the binding pocket followed by partial closure of the entryway. Reversal of these steps enables sterol egress.
- From the Departments of Biochemistry and.
Organizational Affiliation: