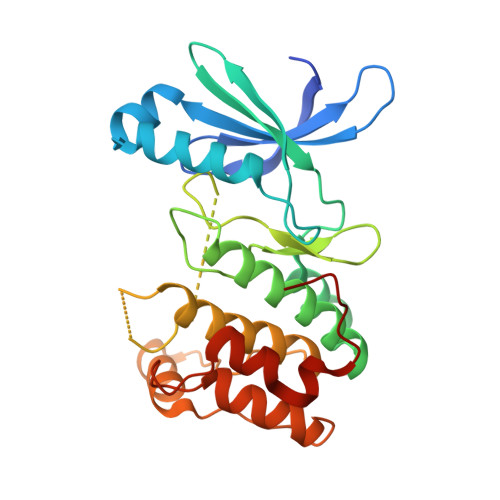
AMP-activated protein kinase selectively inhibited by the type II inhibitor SBI-0206965.
Dite, T.A., Langendorf, C.G., Hoque, A., Galic, S., Rebello, R.J., Ovens, A.J., Lindqvist, L.M., Ngoei, K.R.W., Ling, N.X.Y., Furic, L., Kemp, B.E., Scott, J.W., Oakhill, J.S.(2018) J Biological Chem 293: 8874-8885
- PubMed: 29695504 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA118.003547
- Primary Citation Related Structures:
6BX6 - PubMed Abstract:
Inhibition of the metabolic regulator AMP-activated protein kinase (AMPK) is increasingly being investigated for its therapeutic potential in diseases where AMPK hyperactivity results in poor prognoses, as in established cancers and neurodegeneration. However, AMPK-inhibitory tool compounds are largely limited to compound C, which has a poor selectivity profile. Here we identify the pyrimidine derivative SBI-0206965 as a direct AMPK inhibitor. SBI-0206965 inhibits AMPK with 40-fold greater potency and markedly lower kinase promiscuity than compound C and inhibits cellular AMPK signaling. Biochemical characterization reveals that SBI-0206965 is a mixed-type inhibitor. A co-crystal structure of the AMPK kinase domain/SBI-0206965 complex shows that the drug occupies a pocket that partially overlaps the ATP active site in a type IIb inhibitor manner. SBI-0206965 has utility as a tool compound for investigating physiological roles for AMPK and provides fresh impetus to small-molecule AMPK inhibitor therapeutic development.
- From the Metabolic Signalling Laboratory and.
Organizational Affiliation: