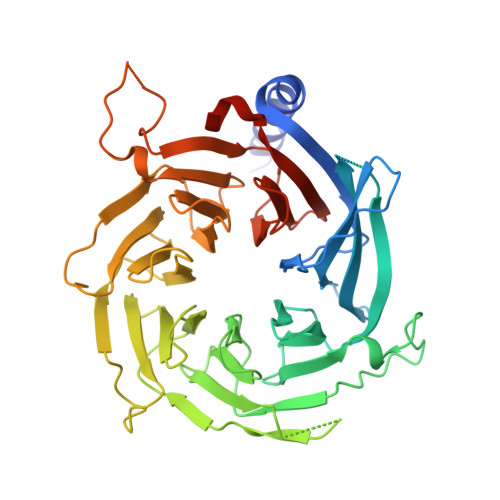
Direct interaction between the PRDM3 and PRDM16 tumor suppressors and the NuRD chromatin remodeling complex.
Ivanochko, D., Halabelian, L., Henderson, E., Savitsky, P., Jain, H., Marcon, E., Duan, S., Hutchinson, A., Seitova, A., Barsyte-Lovejoy, D., Filippakopoulos, P., Greenblatt, J., Lima-Fernandes, E., Arrowsmith, C.H.(2019) Nucleic Acids Res 47: 1225-1238
- PubMed: 30462309 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gky1192
- Primary Citation Related Structures:
6BW3, 6BW4 - PubMed Abstract:
Aberrant isoform expression of chromatin-associated proteins can induce epigenetic programs related to disease. The MDS1 and EVI1 complex locus (MECOM) encodes PRDM3, a protein with an N-terminal PR-SET domain, as well as a shorter isoform, EVI1, lacking the N-terminus containing the PR-SET domain (ΔPR). Imbalanced expression of MECOM isoforms is observed in multiple malignancies, implicating EVI1 as an oncogene, while PRDM3 has been suggested to function as a tumor suppressor through an unknown mechanism. To elucidate functional characteristics of these N-terminal residues, we compared the protein interactomes of the full-length and ΔPR isoforms of PRDM3 and its closely related paralog, PRDM16. Unlike the ΔPR isoforms, both full-length isoforms exhibited a significantly enriched association with components of the NuRD chromatin remodeling complex, especially RBBP4. Typically, RBBP4 facilitates chromatin association of the NuRD complex by binding to histone H3 tails. We show that RBBP4 binds to the N-terminal amino acid residues of PRDM3 and PRDM16, with a dissociation constant of 3.0 μM, as measured by isothermal titration calorimetry. Furthermore, high-resolution X-ray crystal structures of PRDM3 and PRDM16 N-terminal peptides in complex with RBBP4 revealed binding to RBBP4 within the conserved histone H3-binding groove. These data support a mechanism of isoform-specific interaction of PRDM3 and PRDM16 with the NuRD chromatin remodeling complex.
- Department of Medical Biophysics, University of Toronto, Toronto, ON, M5G 1L7, Canada.
Organizational Affiliation: