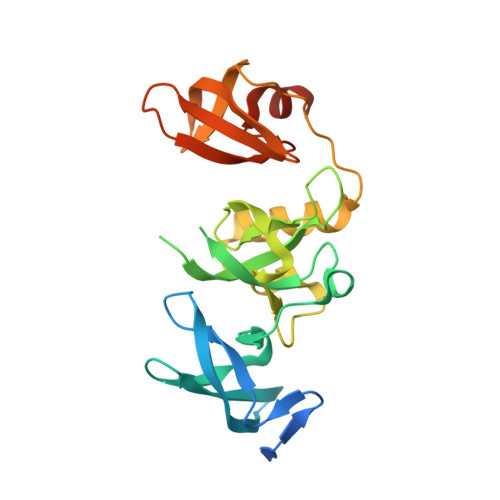
H3K14ac is linked to methylation of H3K9 by the triple Tudor domain of SETDB1.
Jurkowska, R.Z., Qin, S., Kungulovski, G., Tempel, W., Liu, Y., Bashtrykov, P., Stiefelmaier, J., Jurkowski, T.P., Kudithipudi, S., Weirich, S., Tamas, R., Wu, H., Dombrovski, L., Loppnau, P., Reinhardt, R., Min, J., Jeltsch, A.(2017) Nat Commun 8: 2057-2057
- PubMed: 29234025 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-02259-9
- Primary Citation Related Structures:
6BHD, 6BHE, 6BHG, 6BHH, 6BHI - PubMed Abstract:
SETDB1 is an essential H3K9 methyltransferase involved in silencing of retroviruses and gene regulation. We show here that its triple Tudor domain (3TD) specifically binds to doubly modified histone H3 containing K14 acetylation and K9 methylation. Crystal structures of 3TD in complex with H3K14ac/K9me peptides reveal that peptide binding and K14ac recognition occurs at the interface between Tudor domains (TD) TD2 and TD3. Structural and biochemical data demonstrate a pocket switch mechanism in histone code reading, because K9me1 or K9me2 is preferentially recognized by the aromatic cage of TD3, while K9me3 selectively binds to TD2. Mutations in the K14ac/K9me binding sites change the sub-nuclear localization of 3TD. ChIP-seq analyses show that SETDB1 is enriched at H3K9me3 regions and K9me3/K14ac is enriched at SETDB1 binding sites overlapping with LINE elements, suggesting that recruitment of the SETDB1 complex to K14ac/K9me regions has a role in silencing of active genomic regions.
- Department of Biochemistry, Institute of Biochemistry and Technical Biochemistry, Stuttgart University, Allmandring 31, 70569, Stuttgart, Germany.
Organizational Affiliation: