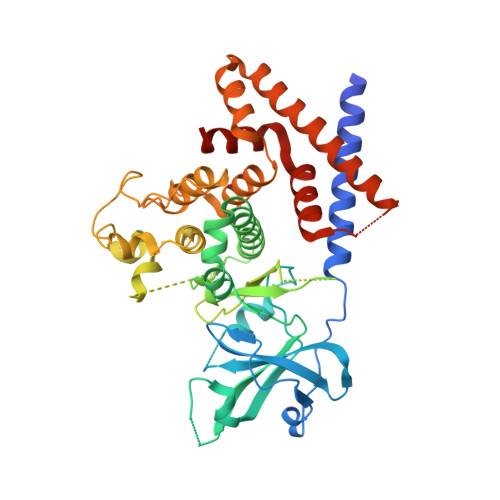
The crystal structure of pseudokinase PEAK1 (Sugen kinase 269) reveals an unusual catalytic cleft and a novel mode of kinase fold dimerization.
Ha, B.H., Boggon, T.J.(2018) J Biological Chem 293: 1642-1650
- PubMed: 29212708 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA117.000751
- Primary Citation Related Structures:
6BHC - PubMed Abstract:
The pseudokinase group encompasses some 10% of protein kinases, but pseudokinases diverge from canonical kinases in key motifs. The two members of the small new kinase family 3 (NKF3) group are considered pseudokinases. These proteins, pseudopodium-enriched atypical kinase 1 (PEAK1, Sugen kinase 269, or SgK269) and pragmin (Sugen kinase 223 or SgK223), act as scaffolds in growth factor signaling pathways, and both contain a kinase fold with degraded kinase motifs at their C termini. These kinases may harbor regions that mediate oligomerization or control other aspects of signal transduction, but a lack of structural information has precluded detailed investigations into their functional roles. In this study, we determined the X-ray crystal structure of the PEAK1 pseudokinase domain to 2.3 Å resolution. The structure revealed that the PEAK1 kinase-like domain contains a closed nucleotide-binding cleft that in this conformation may deleteriously affect nucleotide binding. Moreover, we found that N- and C-terminal extensions create a highly unusual all α-helical split-dimerization region, termed here the split helical dimerization (SHED) region. Sequence conservation analysis suggested that this region facilitates a dimerization mode that is conserved between PEAK1 and pragmin. Finally, we observed structural similarities between the PEAK1 SHED region and the C-terminal extension of the Parkinson's disease-associated kinase PINK1. In summary, PEAK1's kinase cleft is occluded, and its newly identified SHED region may promote an unexpected dimerization mode. Similarities of PEAK1 with the active kinase PINK1 may reclassify the latter as a member of the new kinase family 3 group.
- From the Departments of Pharmacology and.
Organizational Affiliation: