Stable human IgG antibody therapeutics with native framework structure
Christie, M., Rouet, R., Nevoltris, D., Langley, D., Schofield, P., Fabri, L., Tingly, K., Lowe, D., Jermutus, L., Christ, D.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Receptor tyrosine-protein kinase erbB-2 | A [auth C] | 658 | Homo sapiens | Mutation(s): 0 Gene Names: ERBB2, HER2, MLN19, NEU, NGL EC: 2.7.10.1 | 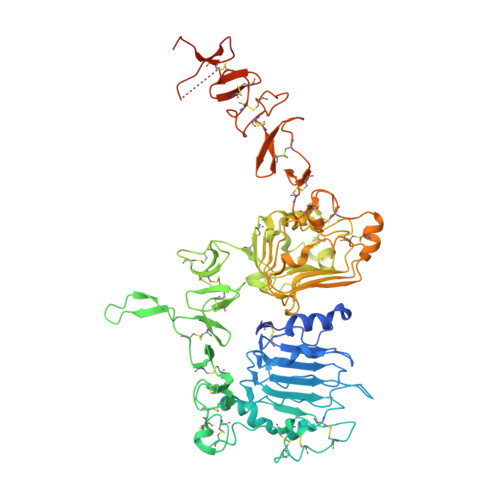 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P04626 GTEx: ENSG00000141736 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P04626 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | Go to GlyGen: P04626-1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Herceptin light chain mutant | B [auth A] | 214 | Homo sapiens | Mutation(s): 0 | 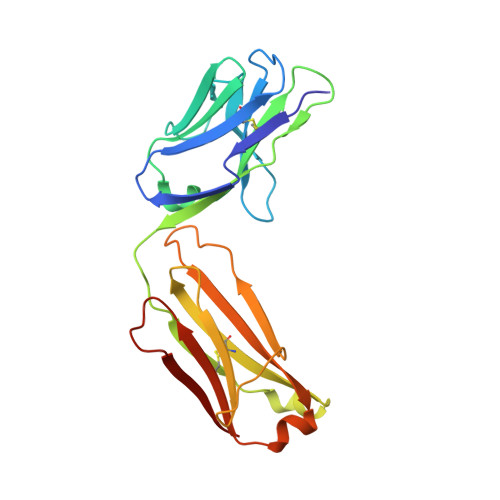 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q7Z3Y4 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Herceptin heavy chain Fab fragment mutant | C [auth B] | 231 | Homo sapiens | Mutation(s): 0 | 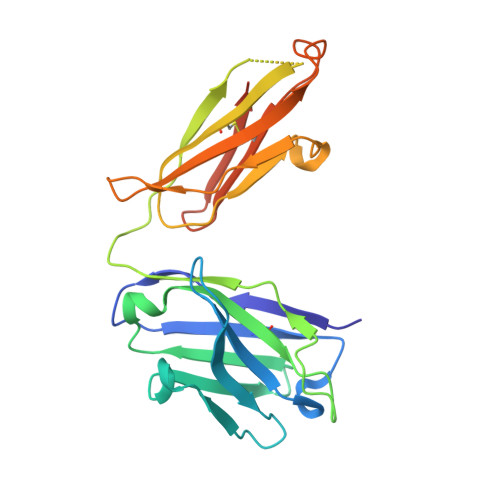 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0DOX5 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG Download:Ideal Coordinates CCD File | D [auth C] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 62.86 | α = 90 |
| b = 114.46 | β = 90 |
| c = 204.279 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| MOSFLM | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Australian Research Council (ARC) | Australia | DE160100608 |
| Australian Research Council (ARC) | Australia | DP160104915 |