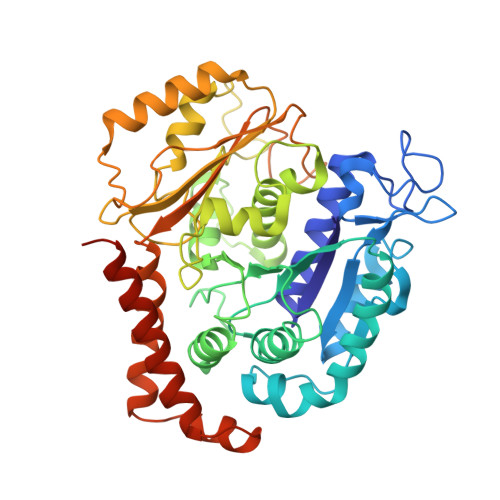
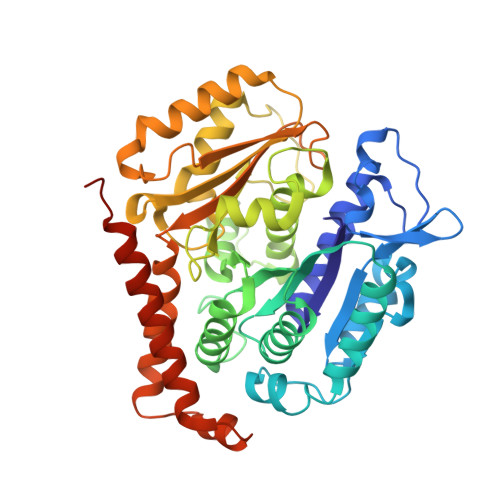
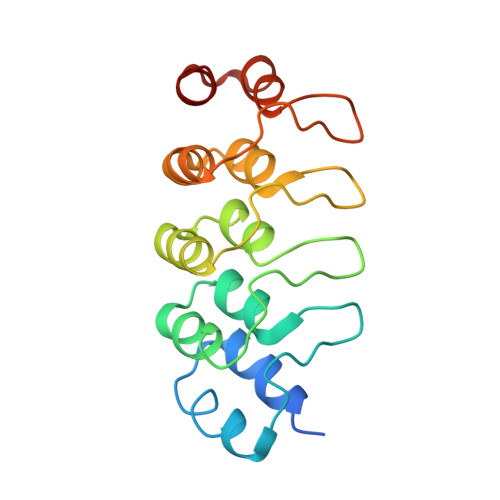
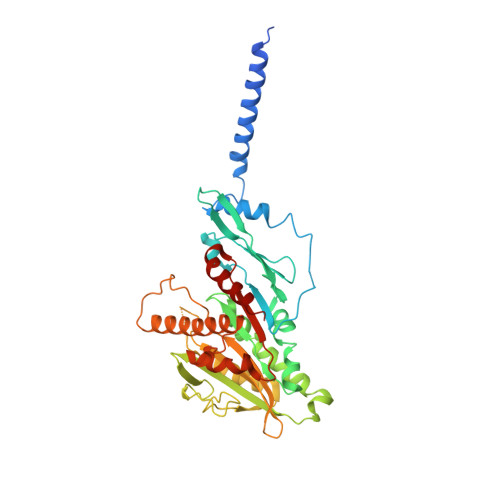
Ternary complex of Kif2A-bound tandem tubulin heterodimers represents a kinesin-13-mediated microtubule depolymerization reaction intermediate.
Trofimova, D., Paydar, M., Zara, A., Talje, L., Kwok, B.H., Allingham, J.S.(2018) Nat Commun 9: 2628-2628
- PubMed: 29980677 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-05025-7
- Primary Citation Related Structures:
6BBN - PubMed Abstract:
Kinesin-13 proteins are major microtubule (MT) regulatory factors that catalyze removal of tubulin subunits from MT ends. The class-specific "neck" and loop 2 regions of these motors are required for MT depolymerization, but their contributing roles are still unresolved because their interactions with MT ends have not been observed directly. Here we report the crystal structure of a catalytically active kinesin-13 monomer (Kif2A) in complex with two bent αβ-tubulin heterodimers in a head-to-tail array, providing a view of these interactions. The neck of Kif2A binds to one tubulin dimer and the motor core to the other, guiding insertion of the KVD motif of loop 2 in between them. AMPPNP-bound Kif2A can form stable complexes with tubulin in solution and trigger MT depolymerization. We also demonstrate the importance of the neck in modulating ATP turnover and catalytic depolymerization of MTs. These results provide mechanistic insights into the catalytic cycles of kinesin-13.
- Department of Biomedical and Molecular Sciences, Queen's University, Kingston, ON, K7L 3N6, Canada.
Organizational Affiliation: