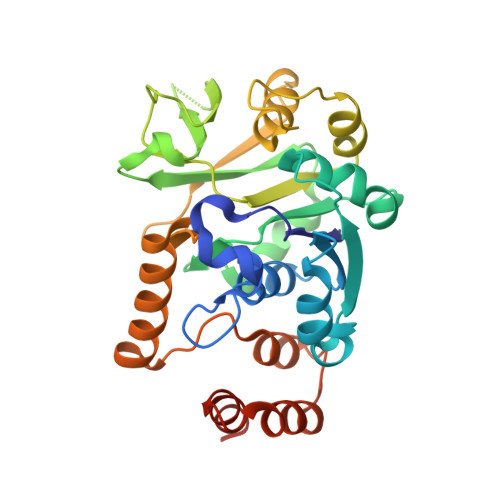
The structure of glucose-1-phosphate thymidylyltransferase from Mycobacterium tuberculosis reveals the location of an essential magnesium ion in the RmlA-type enzymes.
Brown, H.A., Thoden, J.B., Tipton, P.A., Holden, H.M.(2018) Protein Sci 27: 441-450
- PubMed: 29076563 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3333
- Primary Citation Related Structures:
6B5E, 6B5K - PubMed Abstract:
Tuberculosis, caused by the bacterium Mycobacterium tuberculosis, continues to be a major threat to populations worldwide. Whereas the disease is treatable, the drug regimen is arduous at best with the use of four antimicrobials over a six-month period. There is clearly a pressing need for the development of new therapeutics. One potential target for structure-based drug design is the enzyme RmlA, a glucose-1-phosphate thymidylyltransferase. This enzyme catalyzes the first step in the biosynthesis of l-rhamnose, which is a deoxysugar critical for the integrity of the bacterium's cell wall. Here, we report the X-ray structures of M. tuberculosis RmlA in complex with either dTTP or dTDP-glucose to 1.6 Å and 1.85 Å resolution, respectively. In the RmlA/dTTP complex, two magnesium ions were observed binding to the nucleotide, both ligated in octahedral coordination spheres. In the RmlA/dTDP-glucose complex, only a single magnesium ion was observed. Importantly, for RmlA-type enzymes with known three-dimensional structures, not one model shows the position of the magnesium ion bound to the nucleotide-linked sugar. As such, this investigation represents the first direct observation of the manner in which a magnesium ion is coordinated to the RmlA product and thus has important ramifications for structure-based drug design. In the past, molecular modeling procedures have been employed to derive a three-dimensional model of the M. tuberculosis RmlA for drug design. The X-ray structures presented herein provide a superior molecular scaffold for such endeavors in the treatment of one of the world's deadliest diseases.
- Department of Biochemistry, University of Wisconsin, Madison, WI, 53706, USA.
Organizational Affiliation: