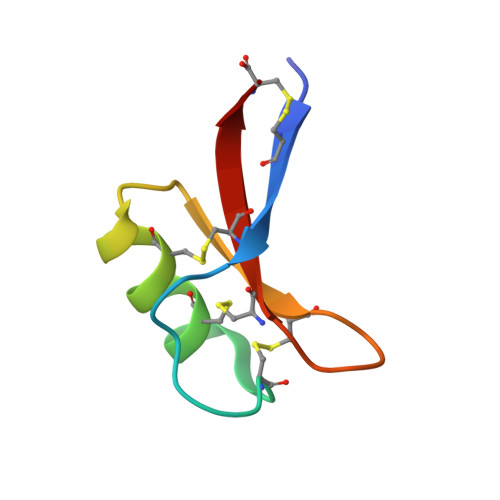
X-ray structure of a carpet-like antimicrobial defensin-phospholipid membrane disruption complex.
Jarva, M., Lay, F.T., Phan, T.K., Humble, C., Poon, I.K.H., Bleackley, M.R., Anderson, M.A., Hulett, M.D., Kvansakul, M.(2018) Nat Commun 9: 1962-1962
- PubMed: 29773800 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-04434-y
- Primary Citation Related Structures:
6B55 - PubMed Abstract:
Defensins are cationic antimicrobial peptides expressed throughout the plant and animal kingdoms as a first line of defense against pathogens. Membrane targeting and disruption is a crucial function of many defensins, however the precise mechanism remains unclear. Certain plant defensins form dimers that specifically bind the membrane phospholipids phosphatidic acid (PA) and phosphatidylinositol 4,5-bisphosphate, thereby triggering the assembly of defensin-lipid oligomers that permeabilize cell membranes. To understand this permeabilization mechanism, here we determine the crystal structure of the plant defensin NaD1 bound to PA. The structure reveals a 20-mer that adopts a concave sheet- or carpet-like topology where NaD1 dimers form one face and PA acyl chains form the other face of the sheet. Furthermore, we show that Arg39 is critical for PA binding, oligomerization and fungal cell killing. These findings identify a putative defensin-phospholipid membrane attack configuration that supports a longstanding proposed carpet mode of membrane disruption.
- Department of Biochemistry and Genetics, La Trobe Institute for Molecular Science, La Trobe University, Melbourne, Victoria, 3086, Australia.
Organizational Affiliation: