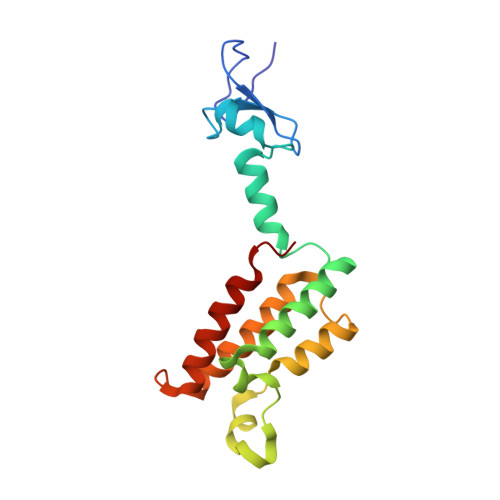
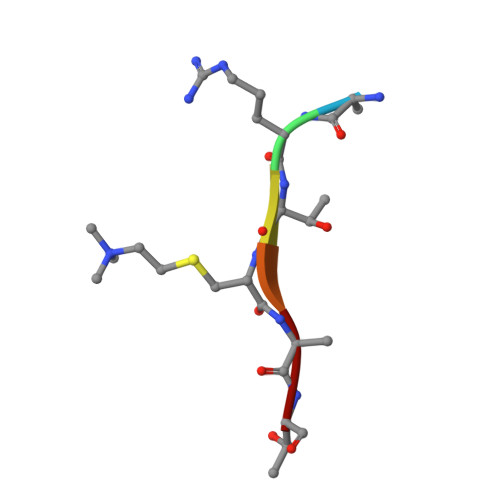
Quantitative and structural assessment of methyllysine analog engagement by cognate binding proteins reveals decrements that can impact qualitative experiment interpretation
Chen, Z., Ruthenburg, A.J., Notti, R.Q., Ueberheide, B., Banaszynski, L.A.To be published.