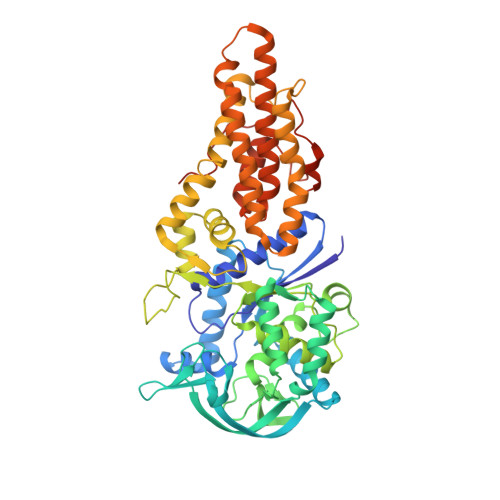
The crystal structure of the drug target Mycobacterium tuberculosis methionyl-tRNA synthetase in complex with a catalytic intermediate.
Barros-Alvarez, X., Turley, S., Ranade, R.M., Gillespie, J.R., Duster, N.A., Verlinde, C.L.M.J., Fan, E., Buckner, F.S., Hol, W.G.J.(2018) Acta Crystallogr F Struct Biol Commun 74: 245-254
- PubMed: 29633973 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X18003151
- Primary Citation Related Structures:
6AX8 - PubMed Abstract:
Mycobacterium tuberculosis is a pathogenic bacterial infectious agent that is responsible for approximately 1.5 million human deaths annually. Current treatment requires the long-term administration of multiple medicines with substantial side effects. Lack of compliance, together with other factors, has resulted in a worrisome increase in resistance. New treatment options are therefore urgently needed. Here, the crystal structure of methionyl-tRNA synthetase (MetRS), an enzyme critical for protein biosynthesis and therefore a drug target, in complex with its catalytic intermediate methionyl adenylate is reported. Phenylalanine 292 of the M. tuberculosis enzyme is in an `out' conformation and barely contacts the adenine ring, in contrast to other MetRS structures where ring stacking occurs between the adenine and a protein side-chain ring in the `in' conformation. A comparison with human cytosolic MetRS reveals substantial differences in the active site as well as regarding the position of the connective peptide subdomain 1 (CP1) near the active site, which bodes well for arriving at selective inhibitors. Comparison with the human mitochondrial enzyme at the amino-acid sequence level suggests that arriving at inhibitors with higher affinity for the mycobacterial enzyme than for the mitochondrial enzyme might be achievable.
- Department of Biochemistry, University of Washington, Seattle, Washington, USA.
Organizational Affiliation: