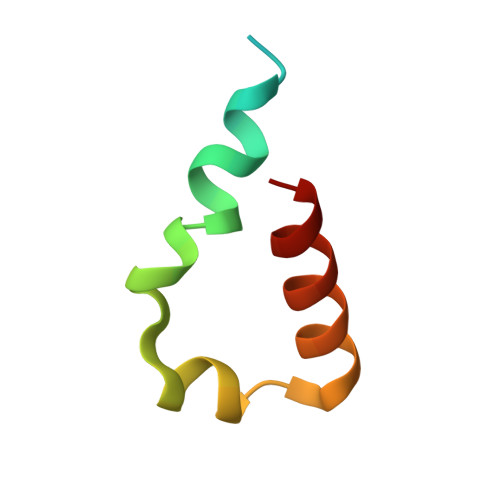
Structures of REV1 UBM2 Domain Complex with Ubiquitin and with a Small-Molecule that Inhibits the REV1 UBM2-Ubiquitin Interaction.
Vanarotti, M., Grace, C.R., Miller, D.J., Actis, M.L., Inoue, A., Evison, B.J., Vaithiyalingam, S., Singh, A.P., McDonald, E.T., Fujii, N.(2018) J Mol Biology 430: 2857-2872
- PubMed: 29864443
- DOI: https://doi.org/10.1016/j.jmb.2018.05.042
- Primary Citation of Related Structures:
6ASR, 6AXD - PubMed Abstract:
REV1 is a DNA damage tolerance protein and encodes two ubiquitin-binding motifs (UBM1 and UBM2) that are essential for REV1 functions in cell survival under DNA-damaging stress. Here we report the first solution and X-ray crystal structures of REV1 UBM2 and its complex with ubiquitin, respectively. Furthermore, we have identified the first small-molecule compound, MLAF50, that directly binds to REV1 UBM2. In the heteronuclear single quantum coherence NMR experiments, peaks of UBM2 but not of UBM1 are significantly shifted by the addition of ubiquitin, which agrees to the observation that REV1 UBM2 but not UBM1 is required for DNA damage tolerance. REV1 UBM2 interacts with hydrophobic residues of ubiquitin such as L8 and L73. NMR data suggest that MLAF50 binds to the same residues of REV1 UBM2 that interact with ubiquitin, indicating that MLAF50 can compete with the REV1 UBM2-ubiquitin interaction orthosterically. Indeed, MLAF50 inhibited the interaction of REV1 UBM2 with ubiquitin and prevented chromatin localization of REV1 induced by cisplatin in U2OS cells. Our results structurally validate REV1 UBM2 as a target of a small-molecule inhibitor and demonstrate a new avenue to targeting ubiquitination-mediated protein interactions with a chemical tool.
- Department of Chemical Biology and Therapeutics, St. Jude Children's Research Hospital, Memphis, TN 38105, USA. Electronic address: murugendra.vanarotti@stjude.org.
Organizational Affiliation: