Crystal structure of Adenosylhomocysteinase from Elizabethkingia anophelis NUHP1 in complex with NAD and Adenosine
Abendroth, J., Delker, S.L., Lorimer, D.D., Edwards, T.E.To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Adenosylhomocysteinase | 445 | Elizabethkingia anophelis NUHP1 | Mutation(s): 0 Gene Names: ahcY, BD94_1936 EC: 3.3.1.1 (PDB Primary Data), 3.13.2.1 (UniProt) | 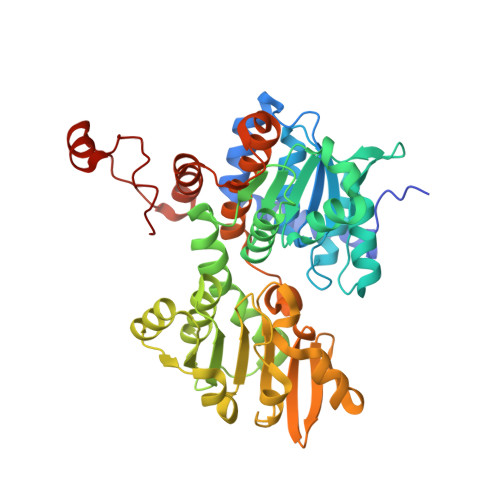 | |
UniProt | |||||
Find proteins for A0A077EDS4 (Elizabethkingia anophelis NUHP1) Explore A0A077EDS4 Go to UniProtKB: A0A077EDS4 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A077EDS4 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAD Download:Ideal Coordinates CCD File | B [auth A] | NICOTINAMIDE-ADENINE-DINUCLEOTIDE C21 H27 N7 O14 P2 BAWFJGJZGIEFAR-NNYOXOHSSA-N |  | ||
| ADN Download:Ideal Coordinates CCD File | C [auth A] | ADENOSINE C10 H13 N5 O4 OIRDTQYFTABQOQ-KQYNXXCUSA-N |  | ||
| MPD Download:Ideal Coordinates CCD File | D [auth A] | (4S)-2-METHYL-2,4-PENTANEDIOL C6 H14 O2 SVTBMSDMJJWYQN-YFKPBYRVSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 132.59 | α = 90 |
| b = 132.59 | β = 90 |
| c = 102.8 | γ = 120 |
| Software Name | Purpose |
|---|---|
| XDS | data reduction |
| XSCALE | data scaling |
| PHASER | phasing |
| PHENIX | refinement |
| ARP | model building |
| Coot | model building |
| PDB_EXTRACT | data extraction |