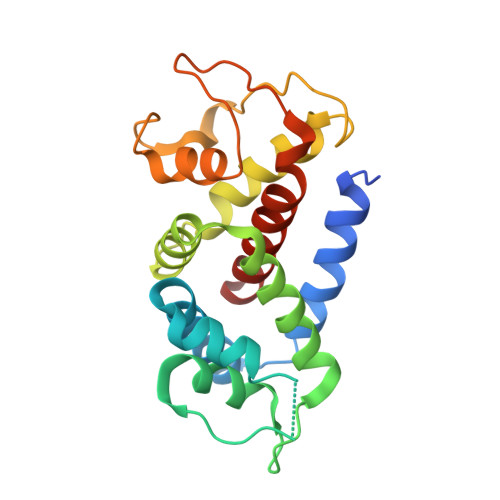
Structures of the Gasdermin D C-Terminal Domains Reveal Mechanisms of Autoinhibition.
Liu, Z., Wang, C., Rathkey, J.K., Yang, J., Dubyak, G.R., Abbott, D.W., Xiao, T.S.(2018) Structure 26: 778-784.e3
- PubMed: 29576317 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2018.03.002
- Primary Citation Related Structures:
6AO3, 6AO4 - PubMed Abstract:
Pyroptosis is an inflammatory form of programmed cell death that plays important roles in immune protection against infections and in inflammatory disorders. Gasdermin D (GSDMD) is an executor of pyroptosis upon cleavage by caspases-1/4/5/11 following canonical and noncanonical inflammasome activation. GSDMD N-terminal domain assembles membrane pores to induce cytolysis, whereas its C-terminal domain inhibits cell death through intramolecular association with the N domain. The molecular mechanisms of autoinhibition for GSDMD are poorly characterized. Here we report the crystal structures of the human and murine GSDMD C-terminal domains, which differ from those of the full-length murine GSDMA3 and the human GSDMB C-terminal domain. Mutations of GSDMD C-domain residues predicted to locate at its interface with the N-domain enhanced pyroptosis. Our results suggest that GSDMDs may employ a distinct mode of intramolecular domain interaction and autoinhibition, which may be relevant to its unique role in pyroptosis downstream of inflammasome activation.
- Department of Pathology, Case Western Reserve University, Cleveland, OH 44106, USA.
Organizational Affiliation: