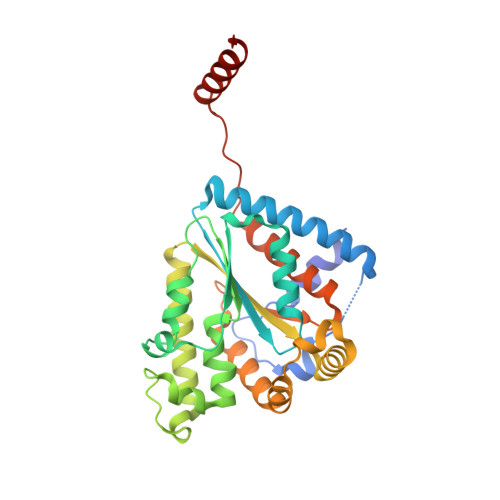
Structural Insights into Substrate Selectivity and Activity of Bacterial Polyphosphate Kinases
Nocek, B., Khusnutdinova, A.N., Ruszkowski, M., Flick, R., Burda, M., Batyrova, K., Brown, G., Mucha, A., Joachimiak, A., Berlicki, L., Yakunin, A.F.(2018) ACS Catal 8: 10746-10760